GMAI-VL & GMAI-VL-5.5M
A General Medical Vision-Language Model Trained on 5.5 Million Image-Text Pairs from 219 Specialised Datasets
Despite remarkable progress in general AI, its application to clinical medicine remains constrained by the absence of domain-specific medical knowledge and the scarcity of large-scale, high-quality multimodal training data. Existing medical datasets are narrow in scope, limited to specific imaging modalities or clinical tasks, and rarely provide the breadth needed to train a truly general model.
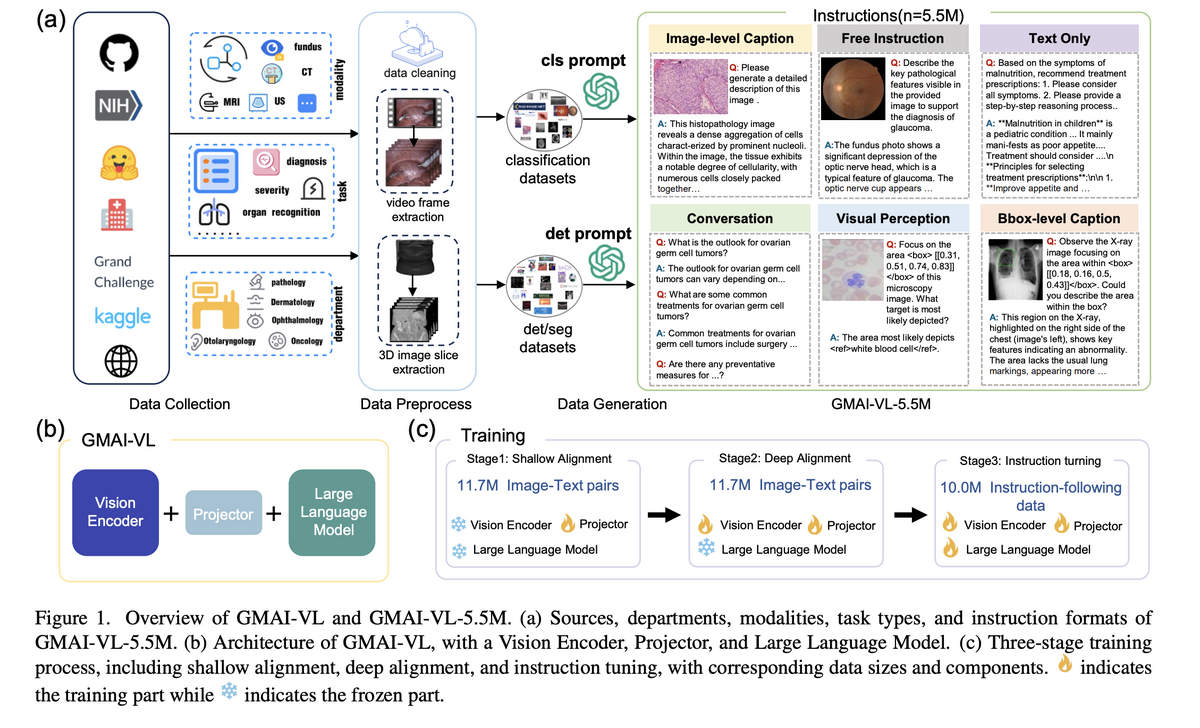
GMAI-VL addresses this challenge end-to-end. We first construct GMAI-VL-5.5M — the largest and most diverse multimodal medical dataset, assembling 5.5 million image-text pairs from 219 specialised medical datasets across 18 clinical departments and 10+ imaging modalities in both English and Chinese — then train GMAI-VL, a state-of-the-art general medical vision-language model, on this foundation using a progressive three-stage training strategy that deepens visual-linguistic alignment at each stage.
🌟 Core Highlights
01 — GMAI-VL-5.5M: Unprecedented Scale & Diversity
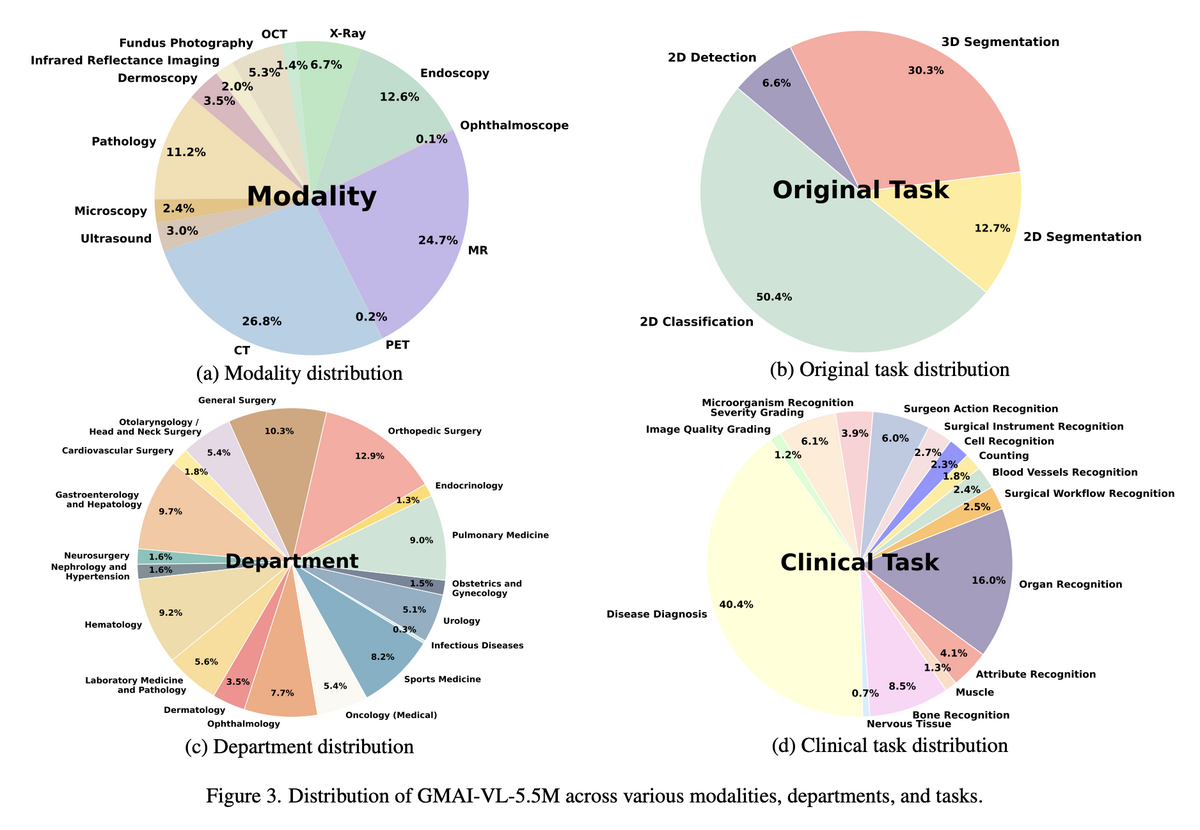
GMAI-VL-5.5M is built by aggregating 219 specialised medical imaging datasets into a unified corpus of 5.5 million high-quality image-text pairs. The dataset spans 18 clinical departments — from radiology and pathology to ophthalmology and endoscopy — and covers 10+ imaging modalities including CT, MRI, X-ray, ultrasound, fundus photography, and microscopy. Critically, all data is fully traceable to source institutions and the dataset is bilingual (EN & CN), making it the most comprehensive foundation for training general medical AI to date.
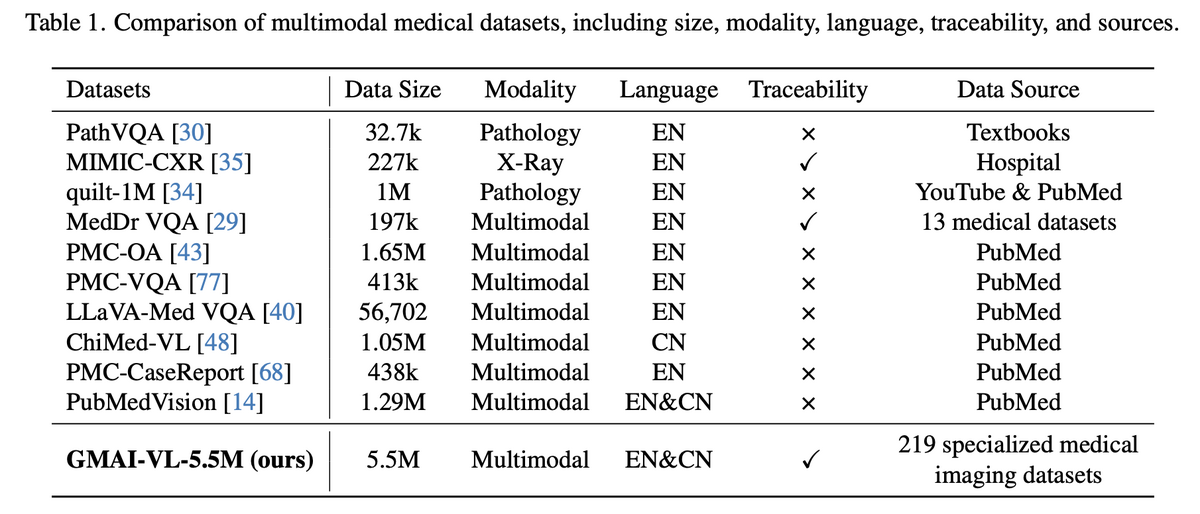
02 — Leading the Medical Multimodal Dataset Landscape
Compared to all prior medical multimodal datasets, GMAI-VL-5.5M stands alone in four critical dimensions: scale (5.5M pairs — far exceeding any competitor), source diversity (219 specialised datasets vs. a handful for others), bilingual support (EN & CN), and full source traceability. This combination makes GMAI-VL-5.5M not just quantitatively larger, but qualitatively superior — enabling generalisation across clinical contexts that no previous dataset could support.
03 — Three-Stage Progressive Training Strategy
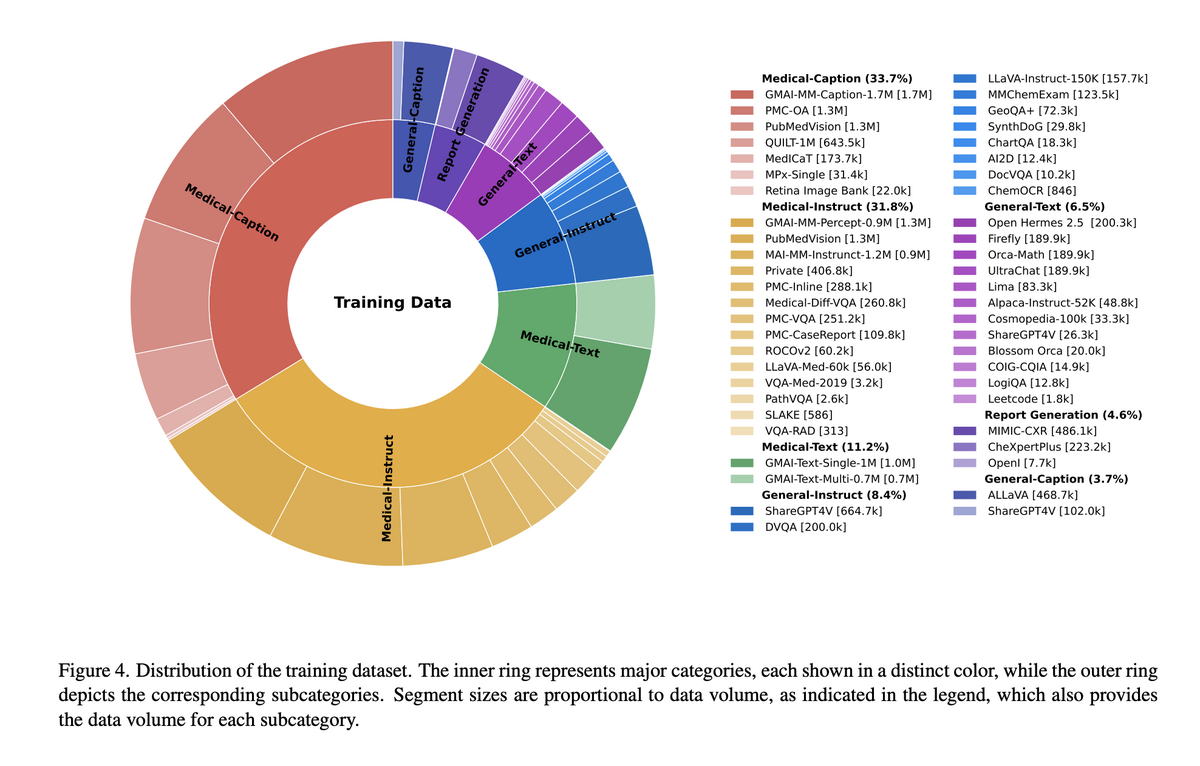
GMAI-VL is trained on a carefully curated corpus spanning medical-caption data (33.7%), medical instruction data (31.8%), general text (13.2%), report generation (4.6%), and general instruction and caption data. Training proceeds in three stages: Stage 1 — Shallow Alignment (11.7M pairs) trains only the projector to align visual and textual representations while keeping the vision encoder and LLM frozen; Stage 2 — Deep Alignment (11.7M pairs) fine-tunes both the projector and LLM for deeper cross-modal fusion; Stage 3 — Instruction Tuning (10.0M pairs) specialises the model for clinical instruction-following and nuanced medical reasoning across diverse task formats.
04 — State-of-the-Art Multimodal Medical Performance
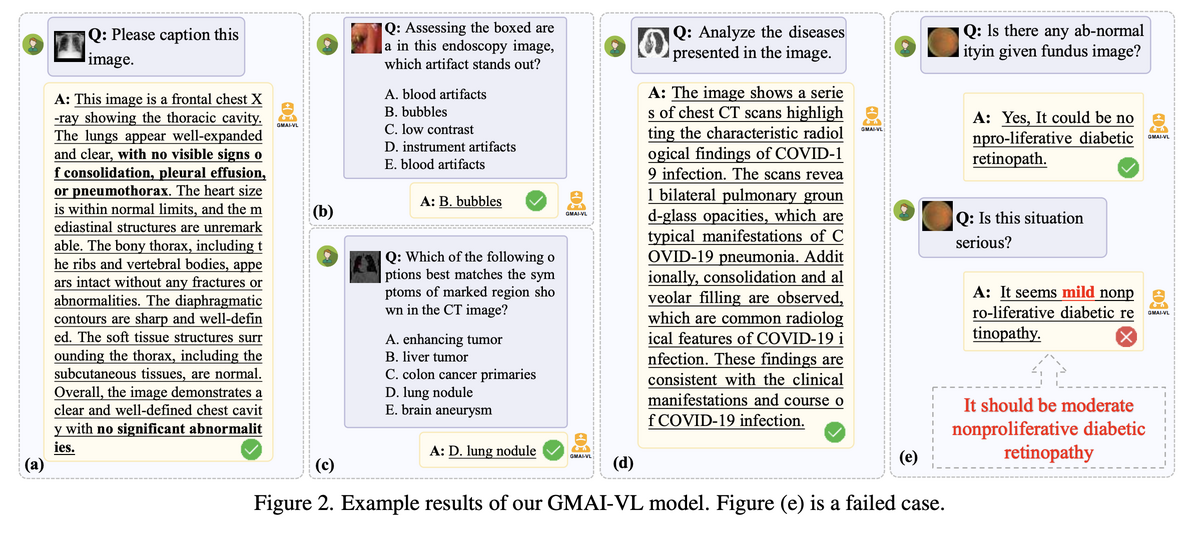
GMAI-VL demonstrates strong performance across a wide range of clinical multimodal tasks: detailed image captioning from chest X-rays and CT scans, visual question answering from endoscopy and fundus images, multi-choice clinical reasoning (identifying artifact types, lesion characteristics, and differential diagnoses), and cross-modal medical inference. The model handles English and Chinese prompts, radiology and pathology images, and both structured (multiple-choice) and open-ended (free-text) instruction formats — validating the generality promised by its training corpus. Experiments confirm state-of-the-art performance across medical VQA and diagnostic reasoning benchmarks.
GMAI-VL establishes a new paradigm for general medical AI by simultaneously solving the twin challenges of data scarcity and model generality. By unifying 219 specialised medical datasets into a single 5.5M-pair corpus and training through a principled three-stage curriculum, GMAI-VL achieves state-of-the-art results on diverse medical VQA and diagnostic reasoning benchmarks. It marks a step-change from narrow, task-specific medical AI towards truly general models capable of assisting clinicians across departments, modalities, and languages.
Key Contributions
- Constructed GMAI-VL-5.5M — the largest and most diverse multimodal medical dataset: 5.5M image-text pairs from 219 specialised datasets, covering 18 clinical departments, 10+ imaging modalities, and bilingual EN & CN with full source traceability.
- Developed GMAI-VL, a state-of-the-art general medical vision-language model, via a three-stage progressive training strategy (shallow alignment → deep alignment → instruction tuning) that systematically strengthens cross-modal clinical reasoning.
- Achieved state-of-the-art performance on multiple medical multimodal VQA and diagnostic reasoning benchmarks, outperforming prior medical-specific and general-purpose vision-language models.
- Fully open-sourced model weights and training code, enabling the community to reproduce, fine-tune, and build upon GMAI-VL for clinical and research applications.
Authors
Tianbin Li, Yanzhou Su, Wei Li, Bin Fu, Zhe Chen, Ziyan Huang, Guoan Wang, Chenglong Ma, Ying Chen, Ming Hu, Yanjun Li, Pengcheng Chen, Xiaowei Hu, Zhongying Deng, Yuanfeng Ji, Jin Ye, Yu Qiao, Junjun He