Scholarly Output
Publications
arXiv · 2025
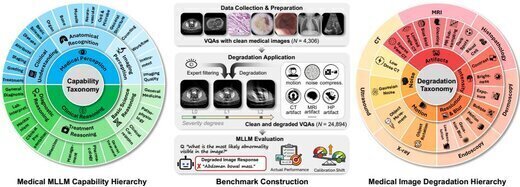
MedQ-Deg: A Multidimensional Benchmark for Evaluating MLLMs Across Medical Image Quality Degradations
We present MedQ-Deg, a comprehensive benchmark for evaluating medical multimodal large language models (MLLMs) under clinically realistic image quality degradations. MedQ-Deg spans 18 degradation types, 30 fine-grained capability dimensions, and 7 imaging modalities with 24,894 question-answer pairs. We introduce the Calibration Shift metric to quantify the gap between model confidence and actual performance, revealing the AI Dunning-Kruger Effect where models maintain inappropriately high confidence despite severe accuracy collapse. Evaluation of 40 mainstream MLLMs shows systematic performance degradation as severity increases.
arXiv · 2025
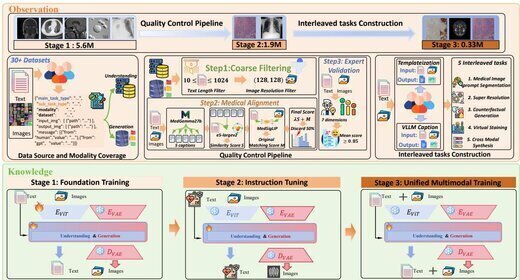
UniMedVL: Unifying Medical Multimodal Understanding and Generation through Observation-Knowledge-Analysis
We propose UniMedVL, the first medical unified multimodal model that combines image understanding and generation within a single architecture without manually reloading model checkpoints. Built on the Observation-Knowledge-Analysis (OKA) framework, we construct UniMed-5M comprising over 5.6M multimodal medical samples and design Progressive Curriculum Learning for systematic capability building. UniMedVL achieves superior performance on 5 medical image understanding benchmarks while matching specialized models in generation quality across 8 imaging modalities, demonstrating that bidirectional knowledge sharing improves both understanding and generation.
arXiv · 2026
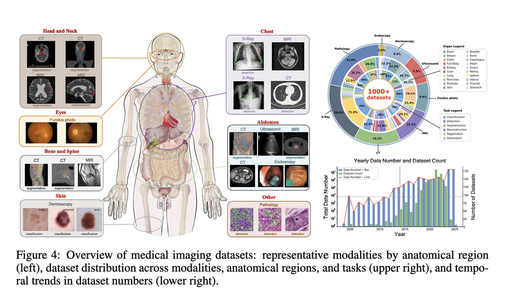
Project Imaging-X: A Survey of 1000+ Open-Access Medical Imaging Datasets for Foundation Model Development
A comprehensive survey and open-source platform integrating 1,000+ open-access medical imaging datasets. We propose a Metadata-Driven Fusion Paradigm (MDFP) to consolidate fragmented small datasets into large, coherent data resources, and build an interactive discovery portal for end-to-end automated dataset integration.
CVPR · 2025
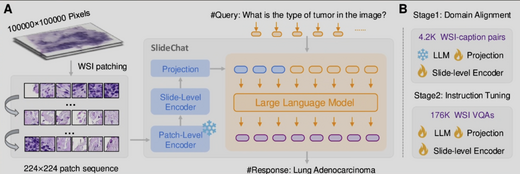
SlideChat: A Large Vision-Language Assistant for Whole-Slide Pathology Image Understanding
We present SlideChat, the first vision-language assistant capable of understanding gigapixel whole-slide images (WSIs). We construct SlideInstruction — a dataset of 4,181 slide-report pairs and 175,753 QA pairs across 13 pathology categories, 20× larger than prior datasets. SlideChat achieves 81.17% accuracy on SlideBench-VQA (TCGA) and surpasses state-of-the-art results on 18 of 22 tasks, establishing a data–model– benchmark framework for next-generation medical vision-language models.
MICCAI · 2025
· Oral
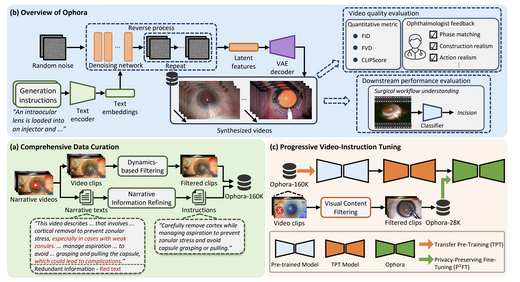
Ophora: A Large-Scale Data-Driven Text-Guided Ophthalmic Surgical Video Generation Model
We propose Ophora, a text-guided surgical video generation model trained on Ophora-160K (160K video-text pairs). A privacy-preserving fine-tuning pipeline using LVLM-based frame filtering produces Ophora-28K. Ophora outperforms existing methods on FID, FVD, and CLIPScore metrics, and synthesized videos significantly improve downstream surgical workflow understanding (e.g., MViTv2 Top-1: 37.92% → 42.24%).
MICCAI · 2025
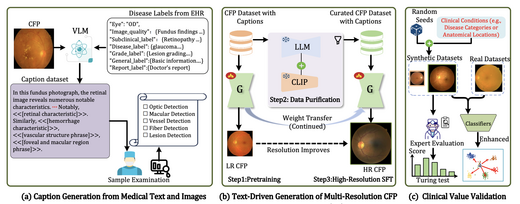
RetinaLogos: Fine-Grained Synthesis of High-Resolution Retinal Images Through Captions
RetinaLogos addresses the chronic shortage of annotated retinal images by generating high-resolution fundus photographs conditioned on free-form natural language. We build RetinaLogos-1400k (14M image-text pairs) and train a three-stage progressive pipeline at 1024×1024 resolution. Expert blind evaluation shows 62.07% of synthetic images are judged as real. DR grading accuracy improves 5–10%, glaucoma detection F1 reaches 0.93.
MICCAI · 2025
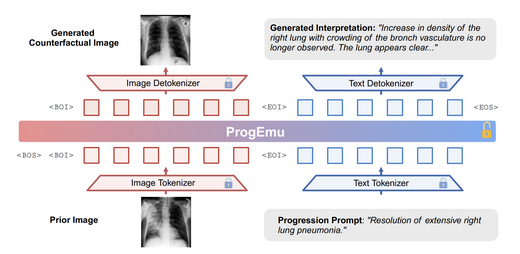
Towards Interpretable Counterfactual Generation via Multimodal Autoregression
We introduce the Interpretable Counterfactual Generation (ICG) task for medical imaging, requiring a model to jointly predict future images and textual radiological explanations given disease progression hypotheses. We build ICG-CXR (11,439 high-quality samples) and propose a multimodal autoregressive model (ProgEmu) that achieves FID 29.21 and ROUGE-L 0.2606, surpassing all existing SOTA methods.
MICCAI · 2025
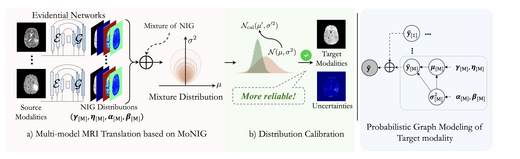
Multi-modal MRI Translation via Evidential Regression and Distribution Calibration
We propose a novel MRI cross-modal translation framework combining Evidential Regression and Distribution Calibration to address missing modality sequences in clinical settings. Each source modality predicts a Normal-Inverse Gamma distribution, fused via MoNIG for uncertainty-aware synthesis. Validated on BraTS2023 and low-field African MRI datasets, achieving PSNR 28.48 dB and UCE 0.089 on cross-center generalization.
CVPR · 2025
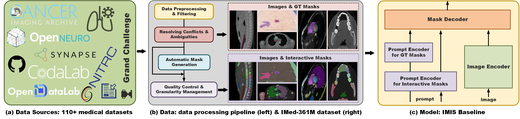
Interactive Medical Image Segmentation: A Benchmark Dataset and Baseline
We introduce IMIS-Bench, a comprehensive benchmark for interactive medical image segmentation with over 361 million masks across diverse imaging modalities. We propose a baseline method and evaluation protocol for interactive segmentation in the medical domain, establishing a standard framework for comparing promptable segmentation approaches in clinical applications.

arXiv · 2025
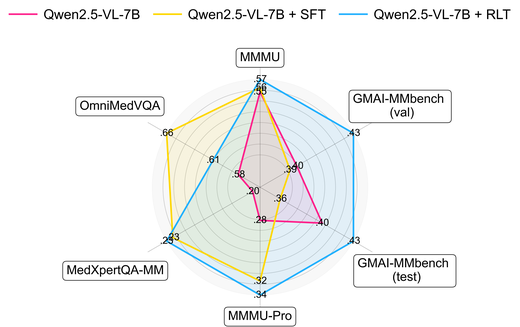
GMAI-VL-R1: Harnessing Reinforcement Learning for Multimodal Medical Reasoning
GMAI-VL-R1 introduces reinforcement learning to medical vision-language models, achieving approximately 30% average accuracy improvement across eight imaging modalities. The model demonstrates that RL-based reasoning can surpass models 36× larger on medical multimodal benchmarks, establishing a new paradigm for efficient medical AI.
arXiv · 2025
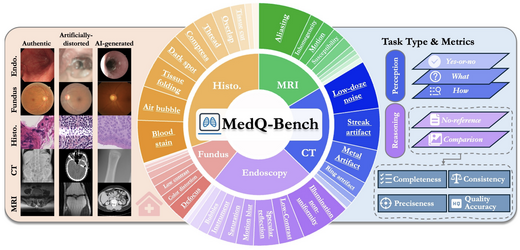
MedQ-Bench: Evaluating and Exploring Medical Image Quality Assessment Abilities in MLLMs
MedQ-Bench introduces a novel perception-reasoning paradigm for medical image quality assessment covering 5 modalities and 40+ quality attributes (3,308 samples). It integrates real clinical images, physical degradation simulations, and AI-synthesized data across three evaluation tracks. Zero-shot evaluation of 14 leading MLLMs shows GPT-4 achieves 68.97% perception accuracy, still 13.5% below expert performance.
arXiv · 2025
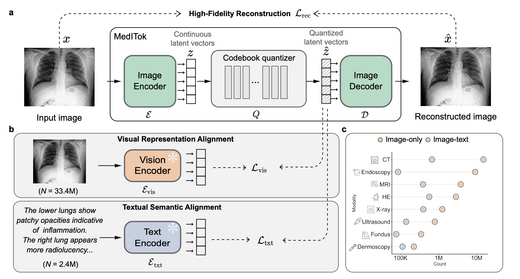
MedITok: A Unified Tokenizer for Medical Image Synthesis and Interpretation
MedITok is the first unified visual tokenizer designed for medical images, addressing the conflict between structural fidelity and semantic richness in joint synthesis and understanding tasks. A two-stage training framework first pre-trains on 30M+ unlabelled images for visual alignment, then fine-tunes on 2M image-text pairs for semantic alignment. MedITok achieves SOTA across reconstruction, classification, generation, and VQA tasks spanning 9 imaging modalities and 30+ datasets.
ICCV · 2025
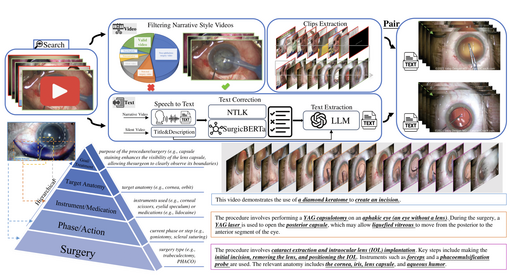
OphCLIP: Hierarchical Retrieval-Augmented Learning for Ophthalmic Surgical Video-Language Pretraining
OphCLIP is a hierarchical retrieval-augmented vision-language pre-training framework designed for understanding ophthalmic surgical workflows. Built on OphVL — a large-scale dataset of 375,000+ hierarchically structured video-text pairs covering surgical phases, instruments, and clinical outcomes — OphCLIP simultaneously learns fine-grained and long-horizon visual representations. Evaluated on 11 datasets for phase recognition and multi-instrument detection, demonstrating robust generalisation and superior performance.
IEEE JBHI · 2026
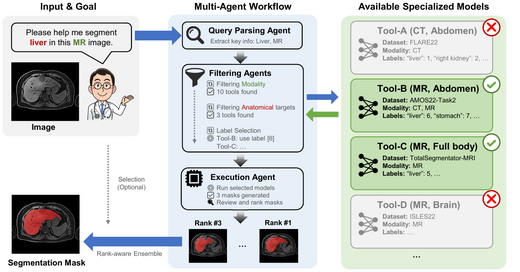
MedSegAgent: A Universal and Scalable Multi-Agent System for Instructive Medical Image Segmentation
MedSegAgent is a multi-agent system for instructive medical image segmentation. Instead of training one universal segmentation model, it orchestrates specialized dataset-specific models through natural language understanding, coarse-to-fine dataset matching, and execution-time result integration. The system integrates 23 datasets and supports 343 segmentation targets across CT, MRI, PET/CT, and ultrasound-related scenarios.
arXiv · 2024
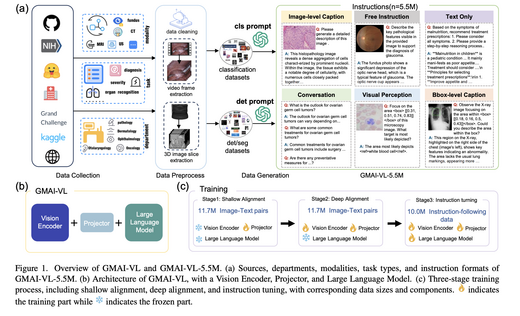
GMAI-VL & GMAI-VL-5.5M: A Large Vision-Language Model and A Comprehensive Multimodal Dataset Towards General Medical AI
GMAI-VL is a general-purpose medical vision-language model trained on GMAI-VL-5.5M — a dataset aggregating hundreds of specialist medical sub-datasets unified into 5.5M high-quality image-text pairs spanning 18 clinical specialties and 10+ imaging modalities. A three-stage training strategy progressively strengthens visual-language alignment and fusion. GMAI-VL achieves or surpasses SOTA on multiple medical multimodal VQA and diagnostic reasoning benchmarks.
NeurIPS · 2024
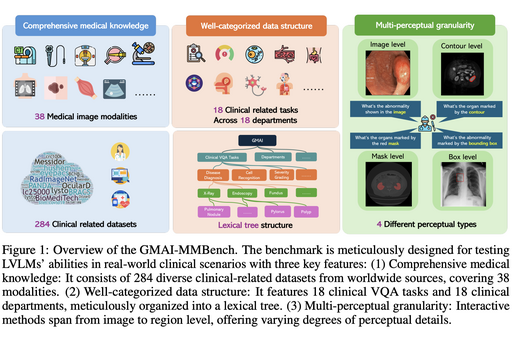
GMAI-MMBench: A Comprehensive Multimodal Evaluation Benchmark Towards General Medical AI
GMAI-MMBench is the most comprehensive general medical AI evaluation platform, covering 284 datasets, 38 imaging modalities, 18 clinical tasks, 18 clinical departments, and 4-level perception granularity with a hierarchical task taxonomy. Evaluation of 50 LVLMs reveals that even top-performing models (e.g., GPT-4o) achieve only 53.96% accuracy, quantifying the challenge of medical multimodal understanding and identifying five common failure modes of current frontier models.

MICCAI · 2024
SAM-Med3D-MoE: Towards a Non-Forgetting Segment Anything Model via Mixture of Experts for 3D Medical Image Segmentation
SAM-Med3D-MoE extends SAM-Med3D with a Mixture-of-Experts architecture to address catastrophic forgetting during task-specific finetuning. By routing different medical imaging tasks to specialized expert modules, the model maintains general segmentation capability while achieving strong task-specific performance across multiple 3D medical imaging benchmarks.
CVPR · 2024
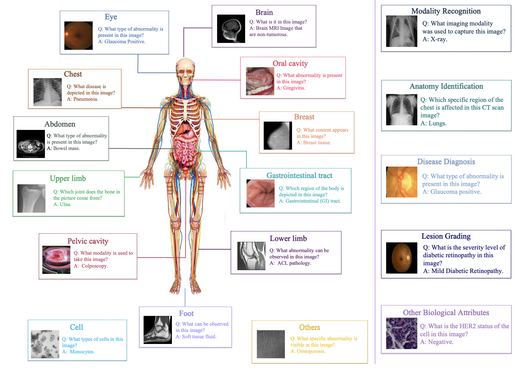
OmniMedVQA: A New Large-Scale Comprehensive Evaluation Benchmark for Medical LVLM
OmniMedVQA is a large-scale medical multimodal VQA benchmark integrating 73 datasets across 12 imaging modalities and 20+ anatomical regions, built from real clinical scenarios. Large-scale experiments reveal two key findings: current LVLMs generally underperform on medical VQA, and surprisingly many medical-specific models lag behind general-purpose models — exposing systematic weaknesses in cross-modal alignment, knowledge generalisation, and robustness.
IEEE TNNLS · 2025
· ECCV 2024 Workshop Oral
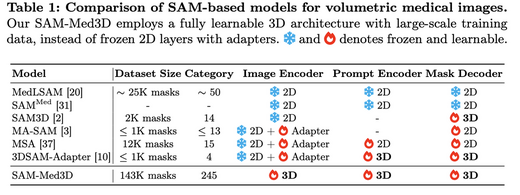
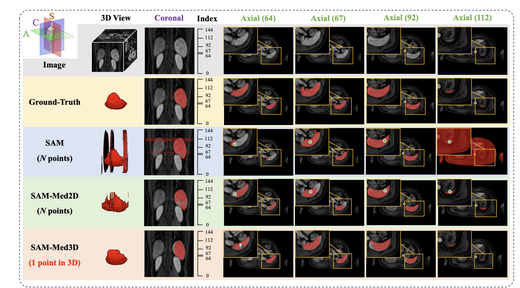
SAM-Med3D: Towards General-purpose Segmentation Models for Volumetric Medical Images
SAM-Med3D adapts the Segment Anything Model (SAM) to 3D volumetric medical images. We construct a large-scale 3D dataset with 21K medical volumes and 131K 3D masks covering 247 categories, then train SAM-Med3D to model three-dimensional context at the voxel level. SAM-Med3D significantly improves segmentation accuracy and robustness over 2D slice-based approaches, providing a universally promptable segmentation backbone for clinical and research applications.
arXiv · 2023
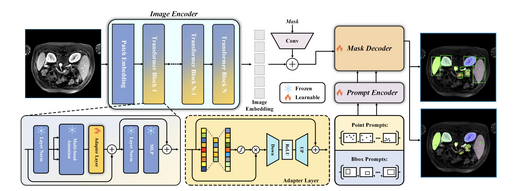
SAM-Med2D
SAM-Med2D adapts the Segment Anything Model (SAM) to 2D medical image segmentation. We construct SA-Med2D-20M — a large-scale benchmark — and train a reliable baseline model that better adapts to medical image structures and distribution shifts. SAM-Med2D significantly improves segmentation accuracy and stability, bridging the gap between natural image segmentation models and clinical medical image analysis.
arXiv · 2023
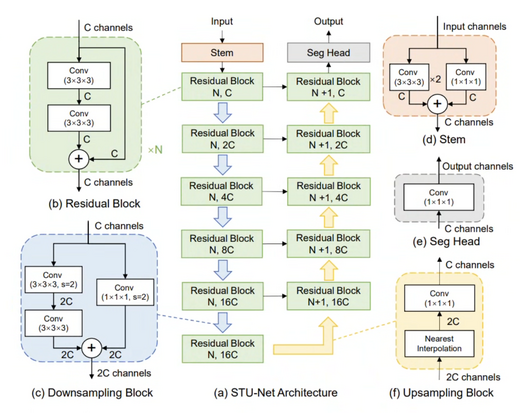
STU-Net: Scalable and Transferable Medical Image Segmentation Models Empowered by Large-Scale Supervised Pre-training
STU-Net is a family of scalable and transferable U-Net models pre-trained on large-scale annotated medical image segmentation datasets. With parameter scales ranging from 14M to 1.4B, STU-Net-H (1.4B parameters) is the largest medical image segmentation model to date. After large-scale supervised pre-training, the models achieve excellent performance on both direct inference and fine-tuning, demonstrating strong scalability and transfer capability.
arXiv · 2025
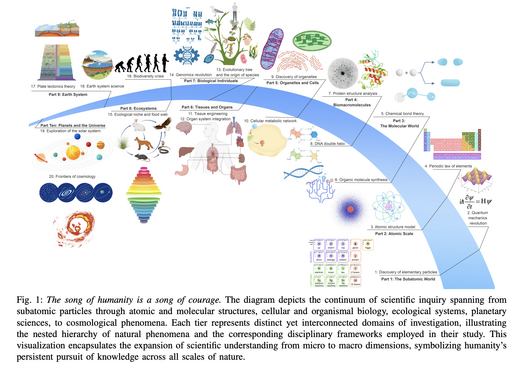
A Survey of Scientific Large Language Models: From Data Foundations to Agent Frontiers
A comprehensive survey of Scientific Large Language Models (Sci-LLMs) produced in collaboration with 20+ leading global institutions, covering 1000+ papers, 600+ key datasets, and SOTA models. The survey systematically reviews the development history, data foundations, model evolution, evaluation frameworks, and agent frontiers of Sci-LLMs, and proposes a roadmap for AI-assisted scientific discovery ecosystems.