Project Imaging-X
A Survey of 1,000+ Open-Access Medical Imaging Datasets for Foundation Model Development
The scarcity of large-scale, diverse, and high-quality training datasets impedes the development of medical imaging foundation models, limiting models to specific tasks, modalities, or anatomical regions. Existing medical imaging datasets are fragmented across narrowly scoped tasks, unevenly distributed across organs and modalities, and lack systematic organisation for broad integration. Prior database surveys often lack detailed statistics, miss recently released large-scale datasets, or provide no systematic framework tailored for foundation model development.
Project Imaging-X addresses the primary obstacle to building such models for healthcare: the scarcity and fragmentation of large-scale medical imaging data. Unlike natural images that can be scraped from the internet by the billions, medical images are difficult to collect due to privacy regulations, the need for specialised equipment, and the high cost of expert clinical annotation.
🌟 Core Highlights
01 — Unprecedented Scale & Systematicity
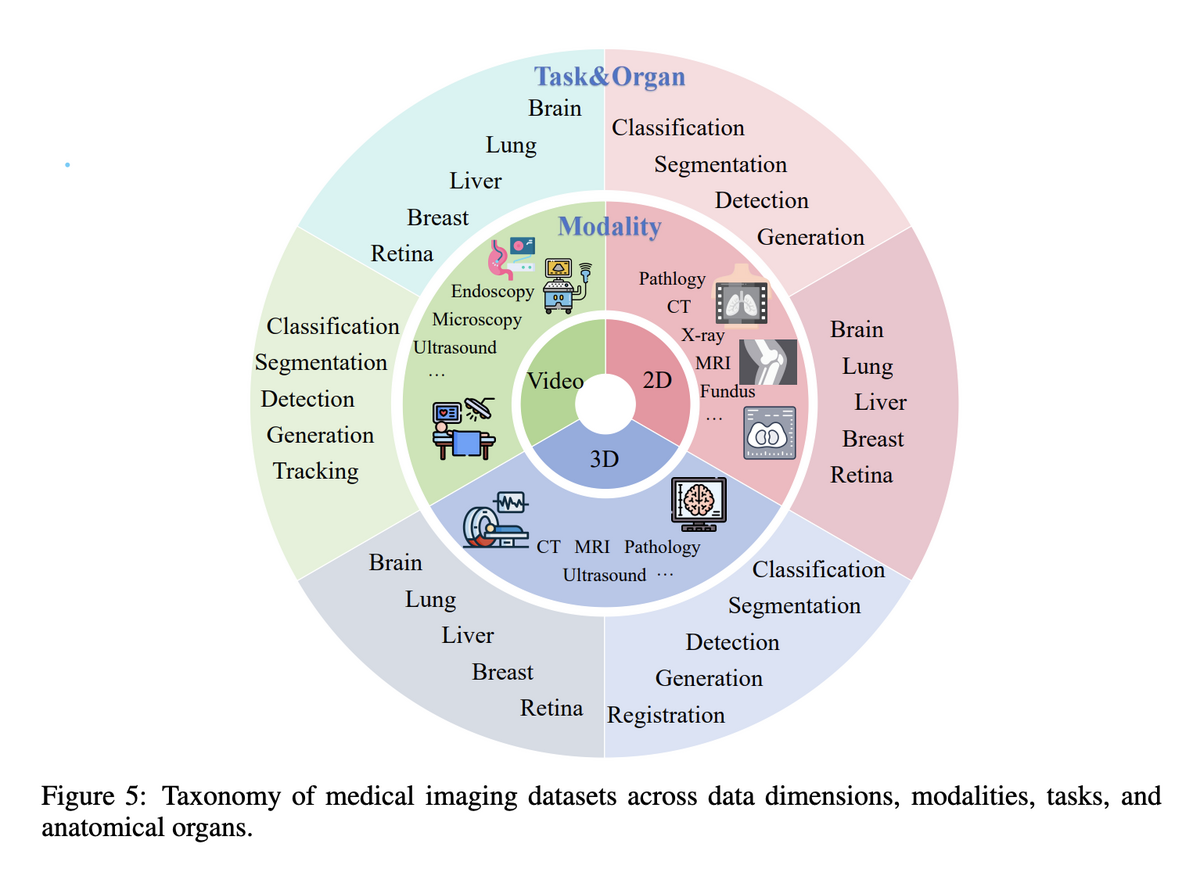
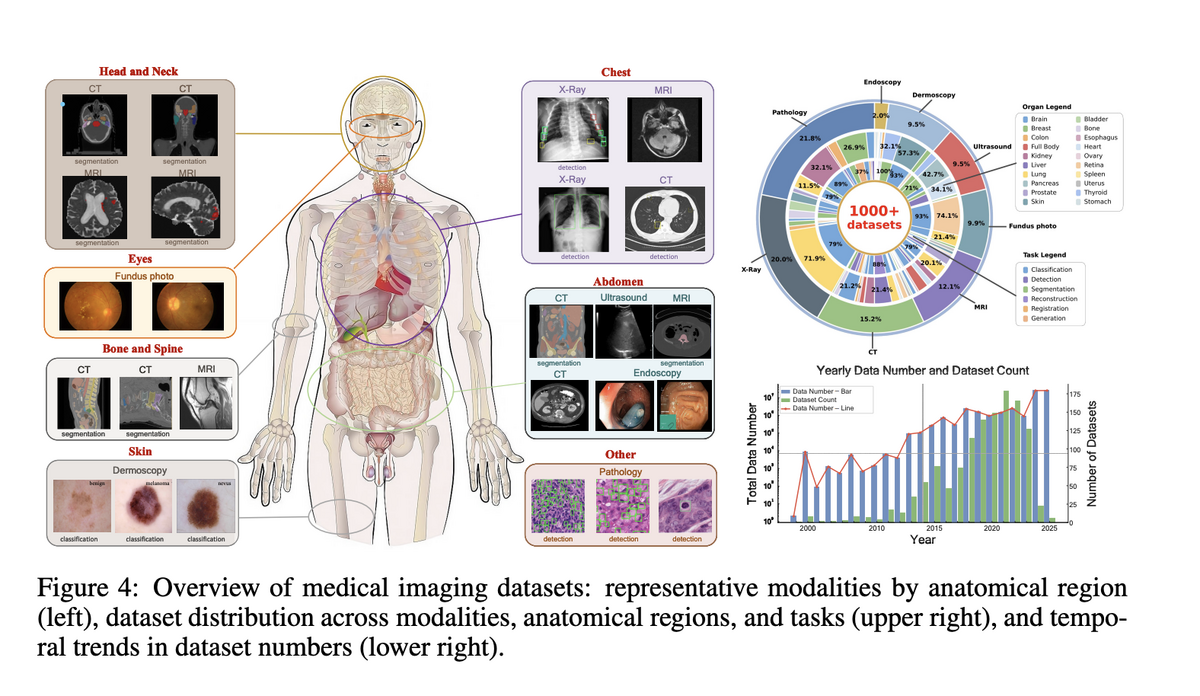
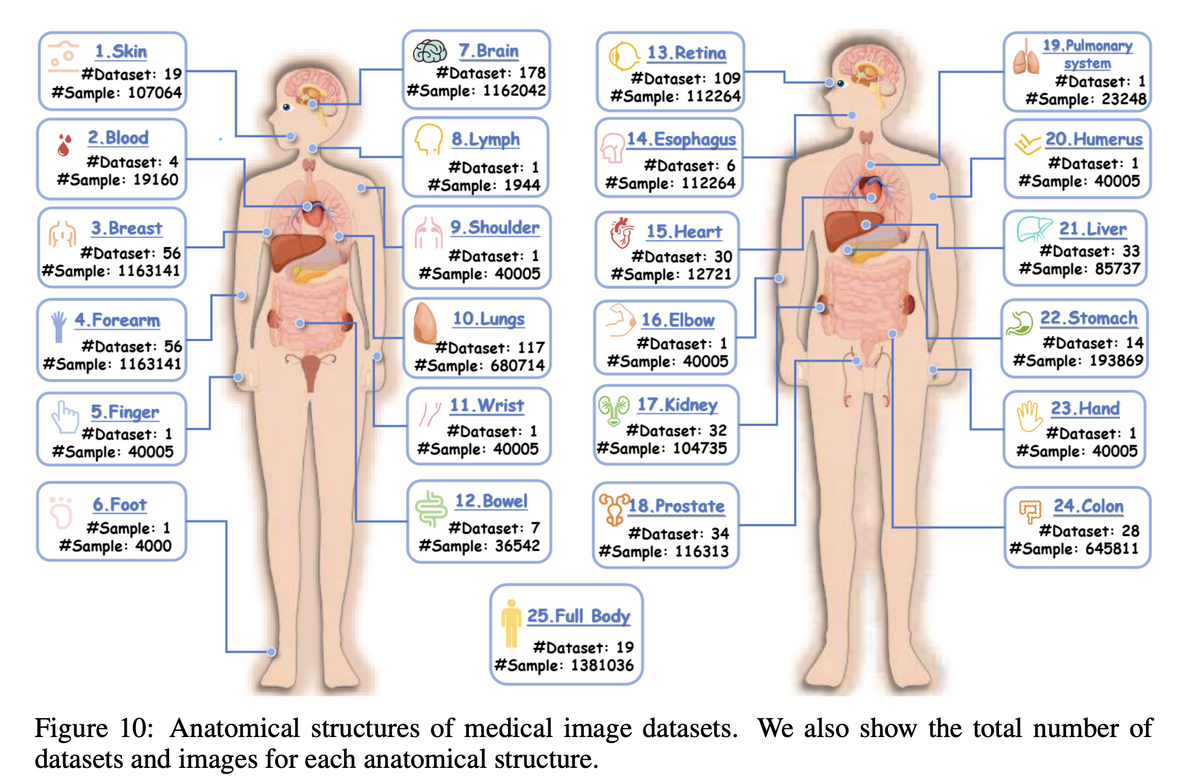
Project Imaging-X is the most comprehensive survey of open-source medical imaging datasets to date, covering 1,000+ datasets across 2D, 3D, and video dimensions, spanning CT, MRI, X-ray, pathology, ultrasound, and more. The survey systematically catalogues task types (classification, segmentation, detection, generation, etc.) and anatomical coverage — providing the community with an accessible, authoritative reference.
02 — Revealing Patterns & Trends in Medical Imaging Data
Through a unified classification framework, the survey provides the first comprehensive analysis of data distribution patterns and distils key findings:
- The core contradiction is not volume but decoupling. Image counts keep rising, but patient-level, 3D-level, longitudinal, and cross-modal coverage has not grown proportionally.
- Distribution reflects acquisition ease, not clinical need. 2D and pathology images dominate partly because they are easier to collect, split, store, and count — "more images" does not equal "more complete clinical information".
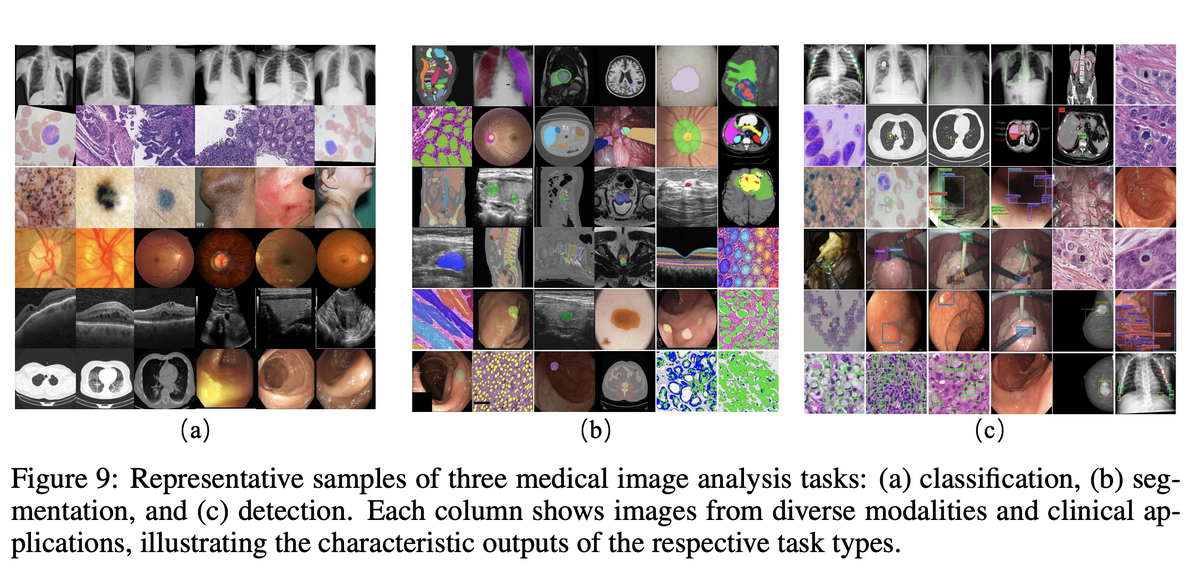
- Task distribution is constrained by annotation cost. Classification and segmentation dominate not because other tasks are less important, but because registration, tracking, VQA, and multimodal reasoning require complex annotations and paired data.
- Recent data expansion is selective. Post-2023 growth concentrates on hot-spot organs (brain, liver, lung, chest) and mainstream modalities, leaving many anatomical regions and rare modalities persistently under-covered.
- The real bottleneck is distribution reconstruction, not more data. Faced with a long-tail, fragmented, imbalanced data ecosystem, naive concatenation fails — what is needed is unified counting, balanced sampling, and principled task design.
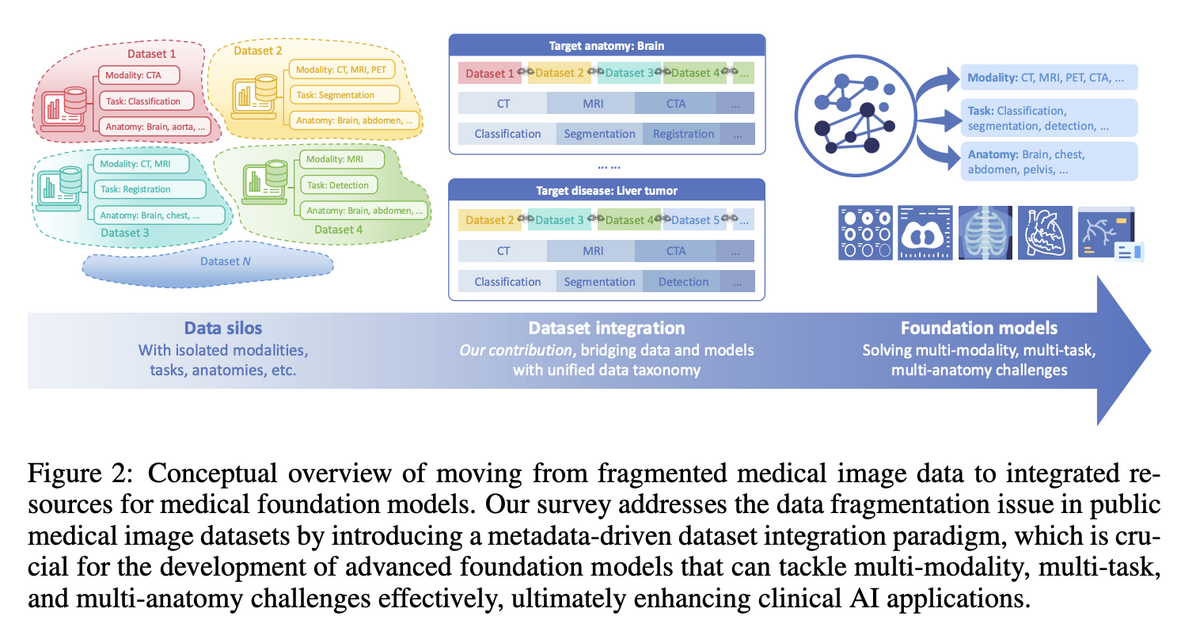
03 — Metadata-Driven Fusion Paradigm (MDFP)
To address fragmentation, the project introduces the Metadata-Driven Fusion Paradigm (MDFP) — a structured methodology for integrating heterogeneous datasets into a coherent corpus through four stages:
Standardise descriptors to authoritative vocabularies (UMLS, MeSH) so "chest", "thorax", and "lung" are recognised as related entities.
Bridge raw ML tasks and clinical meaning by harmonising heterogeneous annotation conventions across datasets.
Group datasets by shared characteristics, assess volume and storage, and flag incompatibilities in imaging protocols or annotation types.
Publish a structured, publicly accessible index enabling fine-grained retrieval — e.g. "all cardiac ultrasound videos with segmentation masks".
04 — Interactive Discovery Portal & Community Tools
Project Imaging-X releases an interactive Medical Dataset Browser — a tool for searching and filtering 1,000+ datasets by modality, anatomy, task, licence, and more. Researchers can instantly find, e.g., all CT datasets with liver and lung annotations under a permissive licence. A companion Python toolkit automates dataset integration, providing fusion blueprints for multi-modal, multi-task foundation model training.
Gap Analysis & Future Directions
The survey identifies critical gaps the community must address to advance medical foundation models:
Organs such as the heart, bowel, and musculoskeletal system have critically insufficient data despite their clinical importance.
Registration, tracking, and multimodal reasoning datasets — critical for clinical intervention — remain far rarer than classification and segmentation data.
Datasets pairing imaging with other clinical data — e.g. radiology + pathology, or images + longitudinal EHR — remain very scarce.
Project Imaging-X represents a systematic effort to map and organise open-access medical imaging data worldwide. By providing a unified taxonomy and metadata-driven integration framework, this work enables the transition from small, task-specific models to large, general-purpose medical foundation models. The identified data gaps serve as a call to action for the clinical community to prioritise data collection in under-represented areas — ensuring that the next generation of medical AI is both powerful and truly comprehensive across all aspects of human health.
Key Contributions
- Catalogued 1,000+ open-access medical imaging datasets with standardised metadata at subject, acquisition, and media level.
- Demonstrated MDFP effectiveness: curated a target-aligned collection of 57 datasets (2.1M+ validated images) for multi-modal, multi-task 2D medical foundation model training.
- Released an interactive web portal ("Medical Dataset Browser") and Python toolkit for efficient dataset discovery, analysis, and integration.
- Provided the first comprehensive gap analysis of open-access medical imaging data, offering clear priority directions for future dataset construction and model training.
Authors
Zhongying Deng, Cheng Tang, Ziyan Huang, Jiashi Lin, Ying Chen, Junzhi Ning, Chenglong Ma, Jiyao Liu, Wei Li, Yinghao Zhu, Shujian Gao, Yanyan Huang, Sibo Ju, Yanzhou Su, Pengcheng Chen, Wenhao Tang, Tianbin Li, Haoyu Wang, Yuanfeng Ji, Hui Sun, Shaobo Min, Liang Peng, Feilong Tang, Haochen Xue, Rulin Zhou, Chaoyang Zhang, Wenjie Li, Shaohao Rui, Weijie Ma, Xingyue Zhao, Yibin Wang, Kun Yuan, Zhaohui Lu, Shujun Wang, Jinjie Wei, Lihao Liu, Dingkang Yang, Lin Wang, Yulong Li, Haolin Yang, Yiqing Shen, Lequan Yu, Xiaowei Hu, Yun Gu, Yicheng Wu, Benyou Wang, Minghui Zhang, Angelica I. Aviles-Rivero, Qi Gao, Hongming Shan, Xiaoyu Ren, Fang Yan, Hongyu Zhou, Haodong Duan, Maosong Cao, Shanshan Wang, Bin Fu, Xiaomeng Li, Zhi Hou, Chunfeng Song, Lei Bai, Yuan Cheng, Yuandong Pu, Xiang Li, Wenhai Wang, Hao Chen, Jiaxin Zhuang, Songyang Zhang, Huiguang He, Mengzhang Li, Bohan Zhuang, Zhian Bai, Rongshan Yu, Liansheng Wang, Yukun Zhou, Xiaosong Wang, Xin Guo, Guanbin Li, Xiangru Lin, Dakai Jin, Mianxin Liu, Wenlong Zhang, Qi Qin, Conghui He, Yuqiang Li, Ye Luo, Nanqing Dong, Jie Xu, Wenqi Shao, Bo Zhang, Qiujuan Yan, Yihao Liu, Jun Ma, Zhi Lu, Yuewen Cao, Zongwei Zhou, Jianming Liang, Shixiang Tang, Qi Duan, Dongzhan Zhou, et al.