MedITok: Unified Medical Image Tokenizer
The First Unified Medical Image Tokenizer for Autoregressive Synthesis and Understanding — Trained on 33M+ Images across 9 Modalities with SOTA on 30+ Benchmarks
Autoregressive modelling has driven major advances in multimodal AI, yet its application to medical imaging remains constrained by the absence of a unified image tokenizer that simultaneously preserves fine-grained anatomical structures and rich clinical semantics across heterogeneous modalities. Existing approaches either optimise for pixel-level reconstruction (e.g., VQGAN) without encoding discriminative features, or capture high-level textual semantics (e.g., CLIP) while failing to retain spatial structures and textures — leaving either synthesis or understanding under-served.
MedITok is the first unified medical image tokenizer that encodes both low-level structural information — supporting faithful image reconstruction and realistic synthesis — and high-level clinical semantics, enabling multimodal medical image comprehension. Built on a principled two-stage training framework that uses visual representation as a bridge, MedITok is trained on over 33 million medical images spanning 9 modalities and 2 million image-text pairs, achieving state-of-the-art performance on 30+ benchmarks across 4 task families: reconstruction, classification, generation, and visual question answering.
Core Highlights
01 — Two-Stage Training: Visual Then Textual Alignment
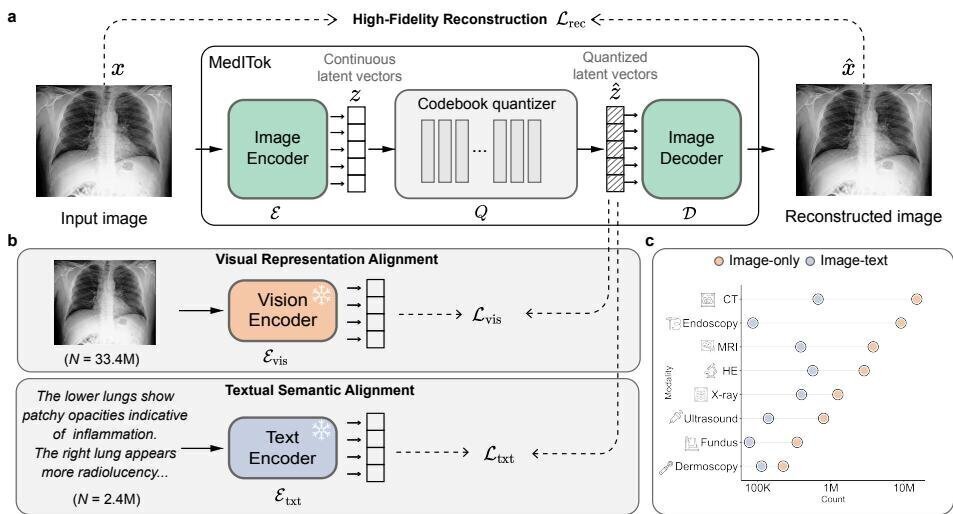
Rather than jointly optimising reconstruction and semantic objectives in a single pass — which risks gradient interference and representation collapse — MedITok introduces a principled two-stage approach. Stage 1 (Visual Representation Alignment) trains the encoder and decoder on 33.4 million unpaired medical images, focusing on reconstruction fidelity with a light semantic constraint from a pretrained vision encoder (BioMed-CLIP). This stage exploits the abundance of unlabelled medical images that existing methods ignore. Stage 2 (Textual Semantic Alignment) refines the encoder on 2.4 million image-text pairs, aligning the learned tokens with fine-grained clinical captions to inject rich semantic information. This progressive strategy avoids the conflicts inherent in naive joint training while building a truly unified latent space.
02 — Unprecedented Scale and Modality Coverage
MedITok is trained on a meticulously curated corpus spanning 9 imaging modalities: CT, dermoscopy, endoscopy, fundus photography, MRI, pathology, ultrasound, X-ray, and OCT. The dataset undergoes rigorous quality control — automated filtering for resolution, intensity range, information content, and clinical relevance, plus manual review to exclude non-clinical content such as tables and plots. This breadth ensures that MedITok learns robust representations across diverse clinical contexts, from chest radiographs to histopathology slides, rather than specialising in a narrow subset of medical imaging.
03 — SOTA across 30+ Benchmarks and 4 Task Families
MedITok achieves rank 1.0 average in reconstruction fidelity (rFID) across 8 modalities despite using a 16× downsampling factor — outperforming tokenizers with only 8× downsampling. Beyond pixel-level metrics, MedITok achieves the highest diagnostic information preservation scores (mAP and AUC) on classification proxy tasks across dermoscopy, fundus, pathology, ultrasound, and X-ray. In linear-probing evaluations for high-level semantic encoding, MedITok consistently outperforms both general-domain and medical-specific tokenizers. When integrated into autoregressive pipelines, MedITok enables competitive medical image synthesis and visual question answering, serving as a scalable foundation component for next-generation multimodal medical models.
MedITok establishes the first unified foundation tokenizer for medical images, demonstrating that a principled two-stage training strategy — leveraging visual representation as a bridge between reconstruction fidelity and semantic richness — can simultaneously excel at low-level encoding, high-level understanding, image synthesis, and visual comprehension. By unlocking the vast pool of unpaired medical images alongside curated image-text pairs, MedITok provides a scalable, modality-agnostic building block for the next generation of autoregressive medical AI models.
Key Contributions
- Proposed a novel two-stage training framework that uses visual representation alignment as a bridge, effectively scaling up with medical image data and progressively building a unified latent space without gradient interference.
- Introduced MedITok, the first Medical Image Tokenizer that unifies the encoding of low-level structural details and high-level clinical semantics within a single model.
- Achieved state-of-the-art performance on over 30 datasets spanning 9 imaging modalities across 4 task families (reconstruction, classification, generation, and VQA), outperforming both general-domain and medical-specific tokenizers.
- Curated a large-scale training corpus of 33M+ medical images and 2M+ image-text pairs with rigorous quality control, with open-source model, code, and data access provided.
Authors
Chenglong Ma, Yuanfeng Ji, Jin Ye, Zilong Li, Chenhui Wang, Junzhi Ning, Wei Li, Lihao Liu, Qiushan Guo, Tianbin Li, Junjun He, Hongming Shan