SAM-Med3D
Towards General-purpose Segmentation Models for Volumetric Medical Images
Medical image segmentation is critical for precise identification of anatomical structures and pathological regions. Current methods are mostly task-specific, requiring specialised models for specific organs, lesions, or imaging modalities. Foundation models like SAM revolutionised 2D segmentation, but applying them to 3D medical volumes poses major challenges — existing approaches either process volumes slice-by-slice (losing spatial context) or use adapter-based methods that cannot fully capture 3D information.
SAM-Med3D addresses these limitations with a fully learnable 3D architecture trained from scratch on SA-Med3D-140K — the largest volumetric medical image segmentation dataset to date, containing 22,000 3D images and 143,000 corresponding 3D masks across 28 imaging modalities, 245+ anatomical target types, and 6 major anatomical categories. The result is a general-purpose model that segments diverse structures and lesions across modalities with just a few 3D prompt points, achieving a 60.12% Dice score improvement over SAM.
🌟 Core Highlights
01 — Fully Native 3D Architecture
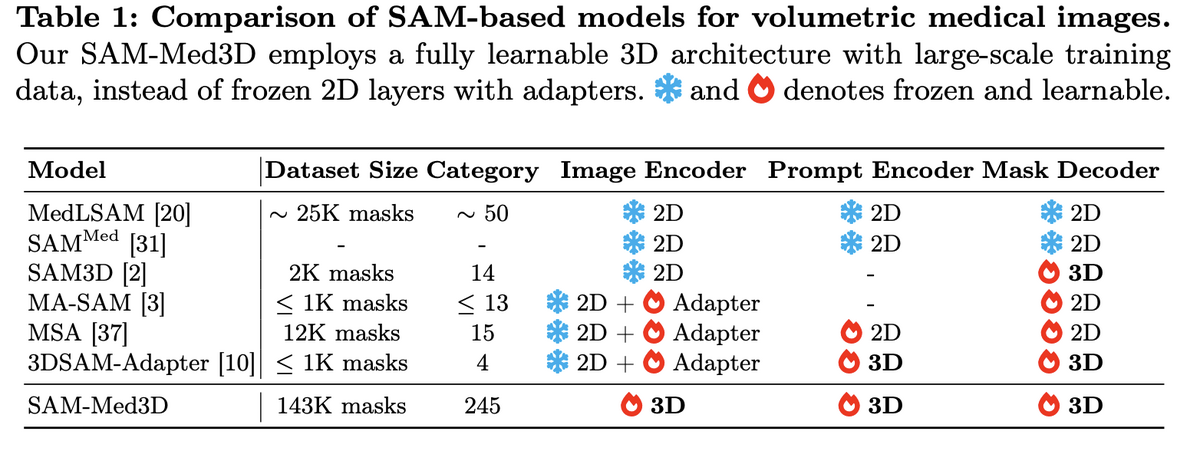
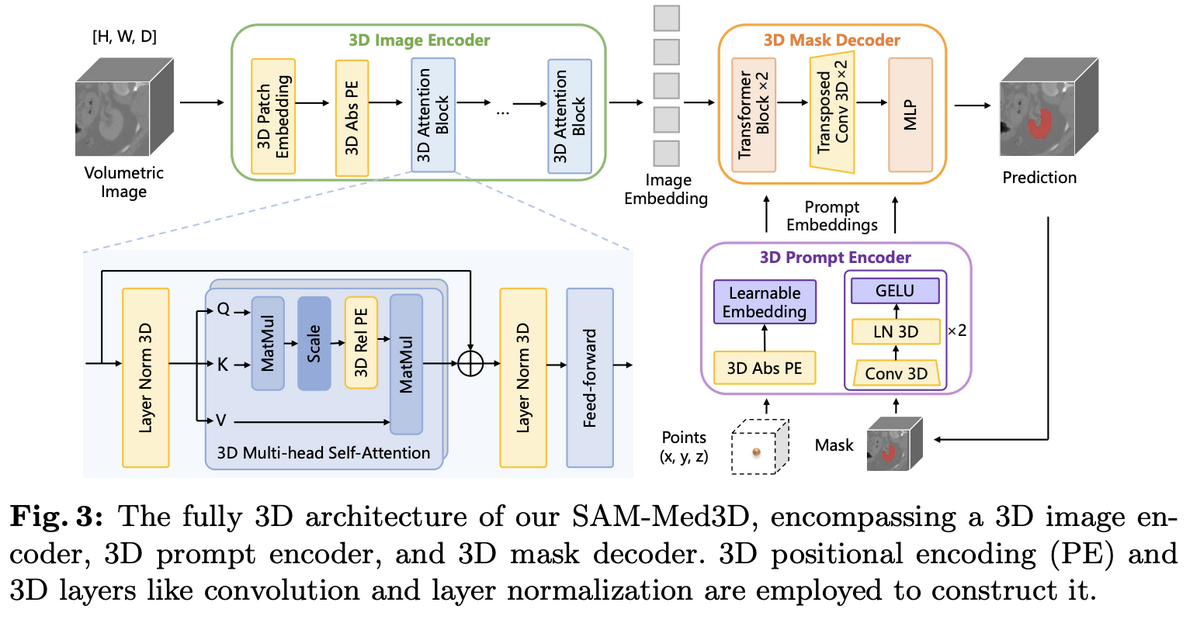
Unlike adapter-based approaches that bolt 3D capability onto frozen 2D backbones, SAM-Med3D is built from the ground up with three fully 3D components: a 3D image encoder, a 3D prompt encoder, and a 3D mask decoder. Initial experiments compared three strategies — 3D adapters on frozen SAM, 2D-to-3D weight transfer with fine-tuning, and training from scratch — and the fully 3D approach clearly won on both seen and unseen targets, avoiding the inherent 2D bias of adapter methods.
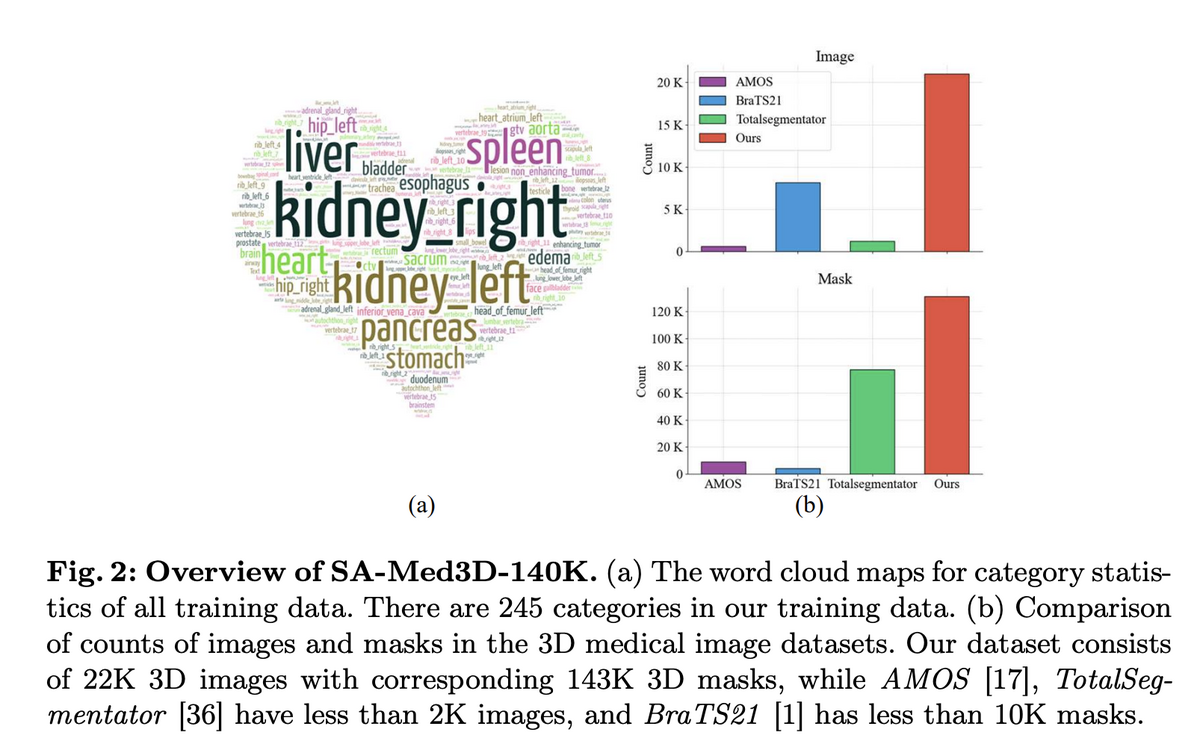
02 — SA-Med3D-140K: Unprecedented Training Scale
The foundation of SAM-Med3D is SA-Med3D-140K, an unprecedented dataset aggregating 70 public and 24 licensed private datasets, spanning CT, ultrasound, and 26 MR sequences across 6 major anatomical categories. A rigorous four-step cleaning pipeline ensures quality: target shape cleaning (removing masks <1cm³), volume size cleaning (excluding >99% background), denoising (eliminating small connected domains), and disambiguation (separating symmetric structures like "kidney" into "left kidney" and "right kidney"). Training uses 131K masks; validation reserves 12 datasets from completely unseen sources plus all ultrasound data to test cross-modal generalisation.
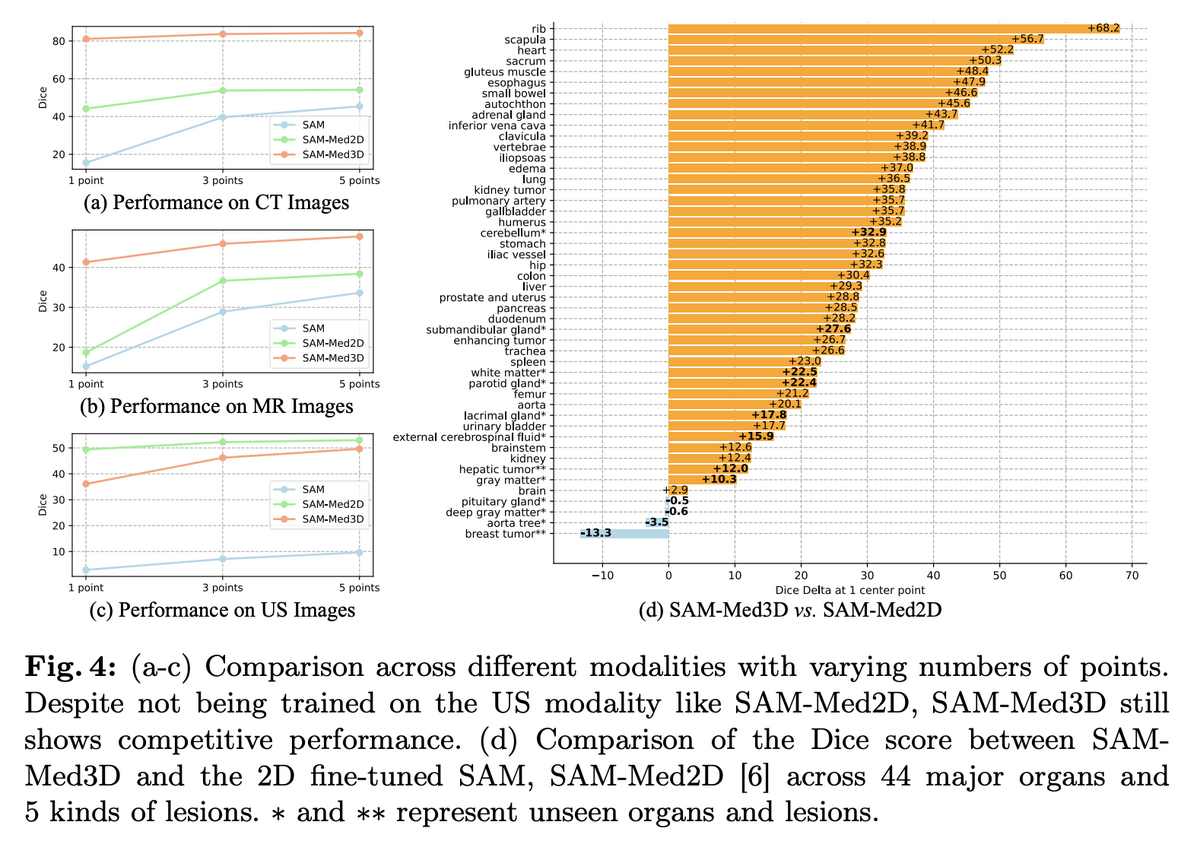
03 — State-of-the-Art Performance Across Modalities
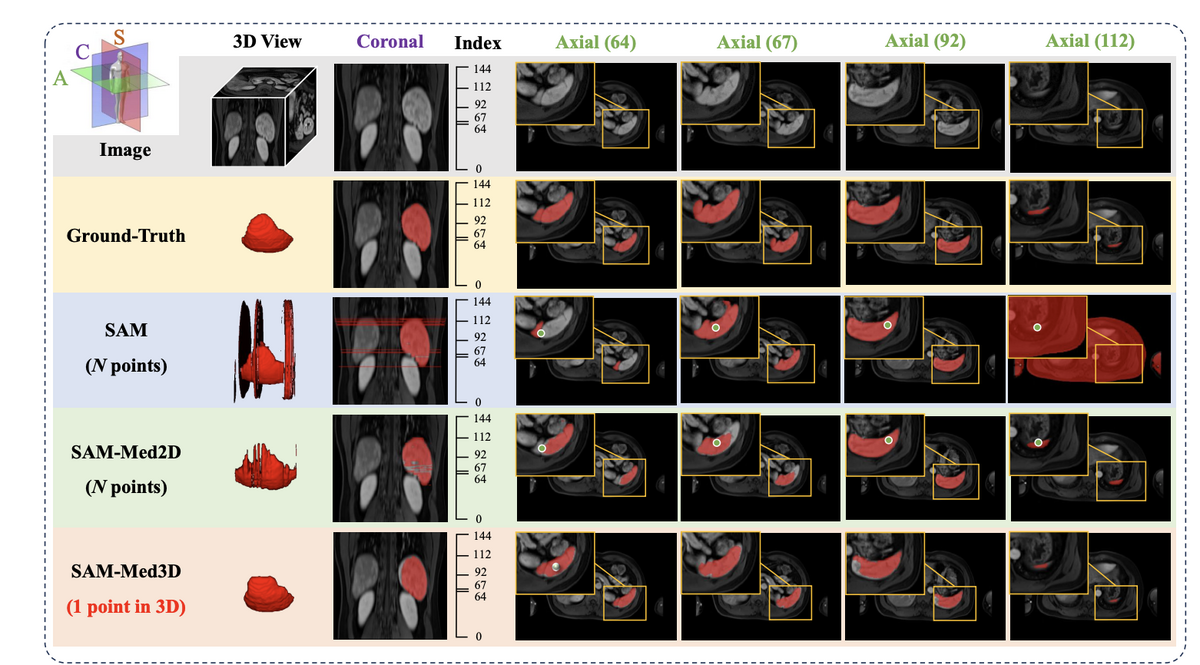
SAM-Med3D achieves 60.12% overall Dice improvement over SAM, with inference time just 1–26% of SAM's depending on target size. Performance is consistent across CT, MR, and ultrasound modalities — notably, despite never being trained on ultrasound data, SAM-Med3D still shows competitive performance on US images. Across 44 major organs and 5 lesion types, SAM-Med3D outperforms SAM-Med2D by up to 68.2% Dice improvement. A two-stage training paradigm (800-epoch pre-training on 131K masks, then fine-tuning on 75K high-quality filtered masks) builds robust general segmentation capability.
04 — Transferability and Clinical Impact
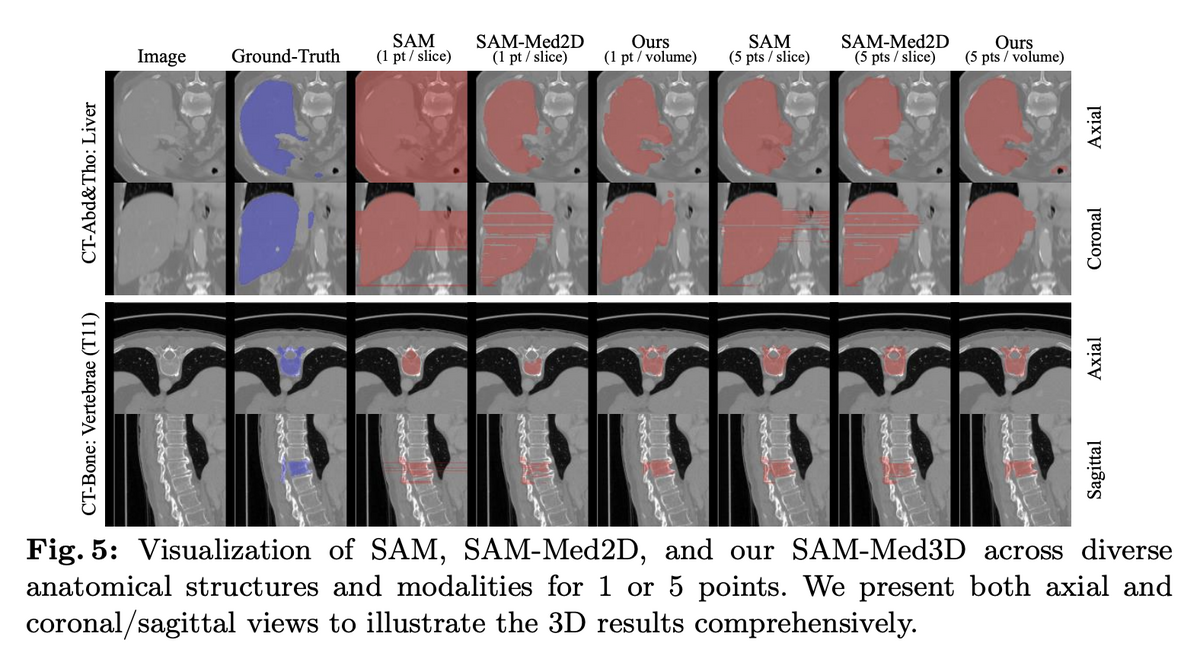
Beyond direct segmentation, SAM-Med3D serves as a powerful pre-trained backbone. When its image encoder is used as a feature extractor for UNETR, downstream semantic segmentation Dice scores improve by up to 5.63% — including on previously unseen challenge datasets. The model requires dramatically fewer prompts than 2D methods (1 point per volume vs. multiple points per slice), with significantly better inter-slice consistency, producing clinically meaningful 3D masks. SAM-Med3D-turbo, a refined version fine-tuned on 44 datasets, further pushes practical performance.
SAM-Med3D demonstrates that a single, fully 3D model can effectively segment diverse anatomical structures and lesions across multiple imaging modalities with minimal user interaction. By training from scratch on the largest volumetric medical dataset (SA-Med3D-140K), the model achieves a paradigm shift from task-specific segmentation to general-purpose medical AI — offering both a powerful standalone tool and a transferable foundation for future research. The open-source release of dataset, code, and models accelerates collective progress toward more universal and accessible medical AI.
Key Contributions
- Introduced a fully native 3D architecture for promptable medical image segmentation, outperforming all adapter-based approaches that bolt 3D onto frozen 2D backbones.
- Created SA-Med3D-140K — 22K 3D images with 143K masks across 28 modalities and 245+ categories — the largest volumetric medical segmentation dataset to date.
- Achieved 60.12% Dice improvement over SAM with inference 1–26× faster, demonstrating strong generalisation across CT, MR, ultrasound, and unseen modalities.
- Open-sourced dataset, code, and model weights (including SAM-Med3D-turbo fine-tuned on 44 datasets) as a reusable foundation for future 3D medical AI research.
Authors
Haoyu Wang, Sizheng Guo, Jin Ye, Zhongying Deng, Junlong Cheng, Tianbin Li, Jianpin Chen, Yanzhou Su, Ziyan Huang, Yiqing Shen, Bin Fu, Shaoting Zhang, Junjun He, Yu Qiao