SlideChat
A Large Vision-Language Assistant for Whole-Slide Pathology Image Understanding
Whole-slide images (WSIs) are the gold standard for digital pathology — gigapixel scans that can exceed 100,000 × 100,000 pixels and encode the full spatial context of tissue samples essential for clinical diagnosis. Yet existing multimodal large language models (MLLMs) are limited to patch-level analysis, discarding the global tissue architecture and cross-region relationships that pathologists rely on. SlideChat is the first vision-language assistant designed specifically to understand these gigapixel slides in their entirety, bridging this critical gap with a scalable, clinically grounded architecture. Accepted at CVPR 2025.
To power SlideChat's development, the team created SlideInstruction — the largest instruction-following dataset for WSIs, containing 4.2K WSI captions and 176K VQA pairs derived from 4,915 TCGA pathology reports using GPT-4-guided curation. They further established SlideBench, a rigorous multi-modal evaluation benchmark covering captioning and visual question-answering across multiple cancer types, validated by expert pathologists. SlideChat achieves state-of-the-art performance on 18 of 22 tasks.
🌟 Core Highlights
01 — SlideInstruction: Largest WSI Instruction-Following Dataset
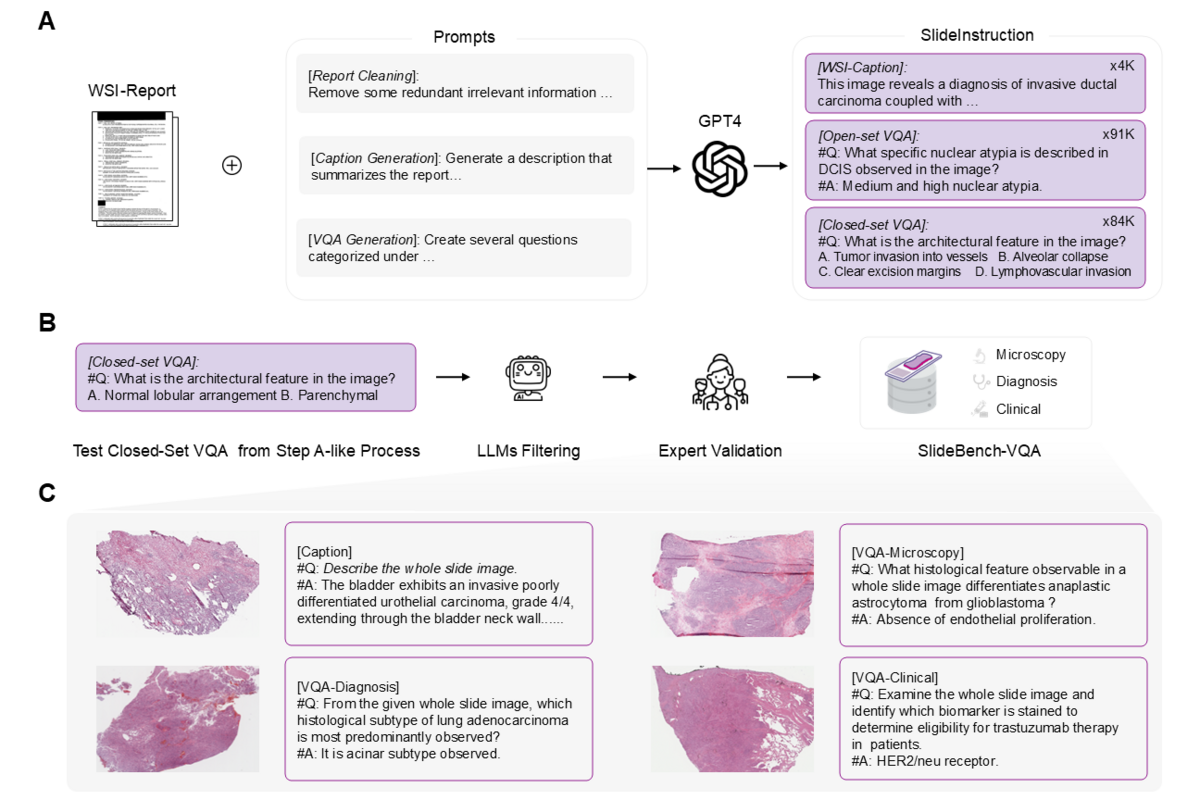
The team curated SlideInstruction from 4,915 WSI-report pairs in the TCGA database, covering 10 cancer types across 4,028 patients. GPT-4 was used to extract high-quality instruction data in three stages: report refinement to remove administrative noise, caption generation producing 4,181 concise clinical WSI descriptions, and VQA pair generation yielding 175,753 question-answer pairs. These span 13 narrow categories within three broad clinical domains — microscopy, diagnosis, and clinical considerations — mirroring the real pathology workflow.
02 — Novel Four-Component Architecture for Gigapixel Understanding
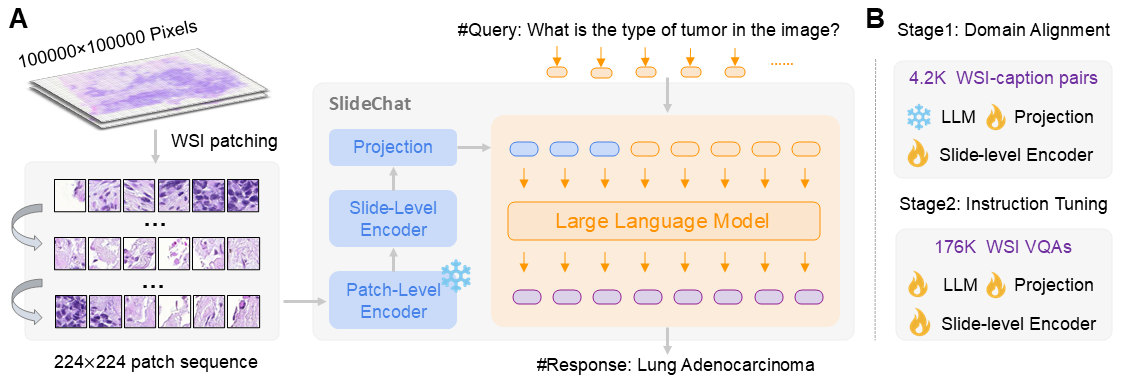
SlideChat serializes each WSI into non-overlapping 224×224 patches at 20× magnification. A frozen CONCH patch-level encoder extracts fine-grained local features (cellular structures, nuclear morphology). A LongNet-based slide-level encoder with sparse attention then processes the full patch token sequence — handling sequences far longer than standard transformers — to generate contextual slide-level embeddings that capture global tissue architecture. A multimodal projector aligns visual features with the language model's embedding space, and Qwen2.5-7B-Instruct serves as the backbone LLM. Training uses two stages: Cross-Domain Alignment (projector + slide encoder only, using 4.2K captions), then Visual Instruction Learning (all components trainable, using 176K VQA pairs).
03 — Benchmark Results: 81.17% Accuracy, SOTA on 18 of 22 Tasks
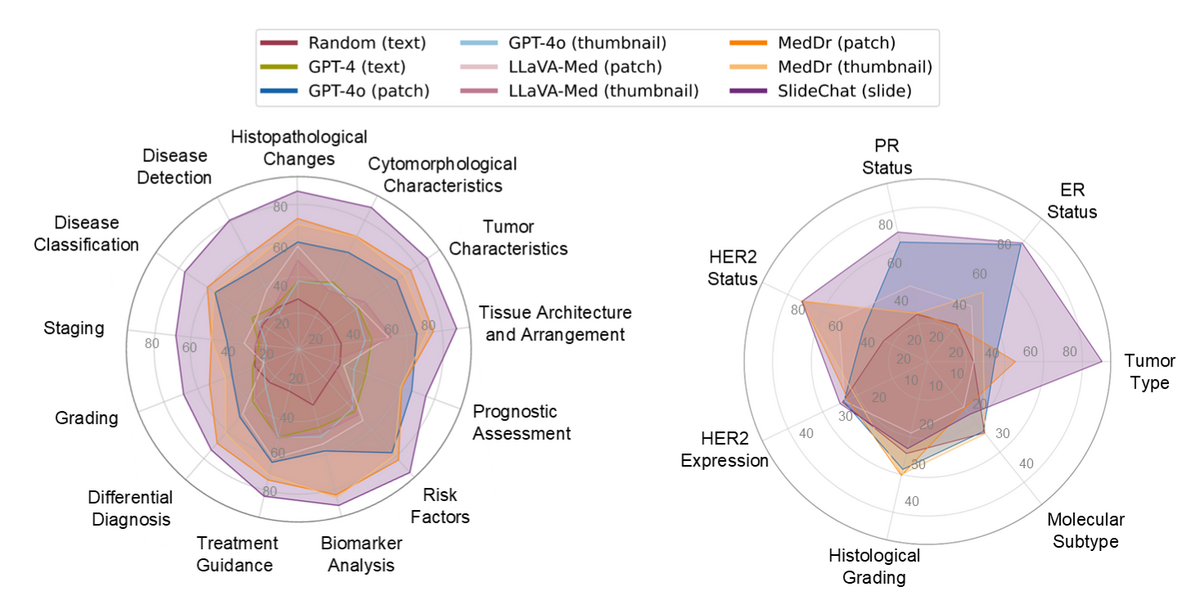
SlideChat achieves 81.17% overall accuracy on SlideBench-VQA (TCGA) — a 13.47% improvement over the second-best model — and 54.14% on the out-of-distribution SlideBench-VQA (BCNB) in zero-shot settings. Compared to general MLLMs (GPT-4o patch: 57.91%; thumbnail: 34.07%) and specialized models (MedDr: 67.70%), SlideChat's holistic WSI understanding enables performance that patch-based approaches cannot match. Results hold across all three clinical domains: microscopy (87.64%), diagnosis (73.27%), and clinical tasks (84.26%).
04 — Clinical Conversational AI: Multi-Turn Diagnostic Reasoning
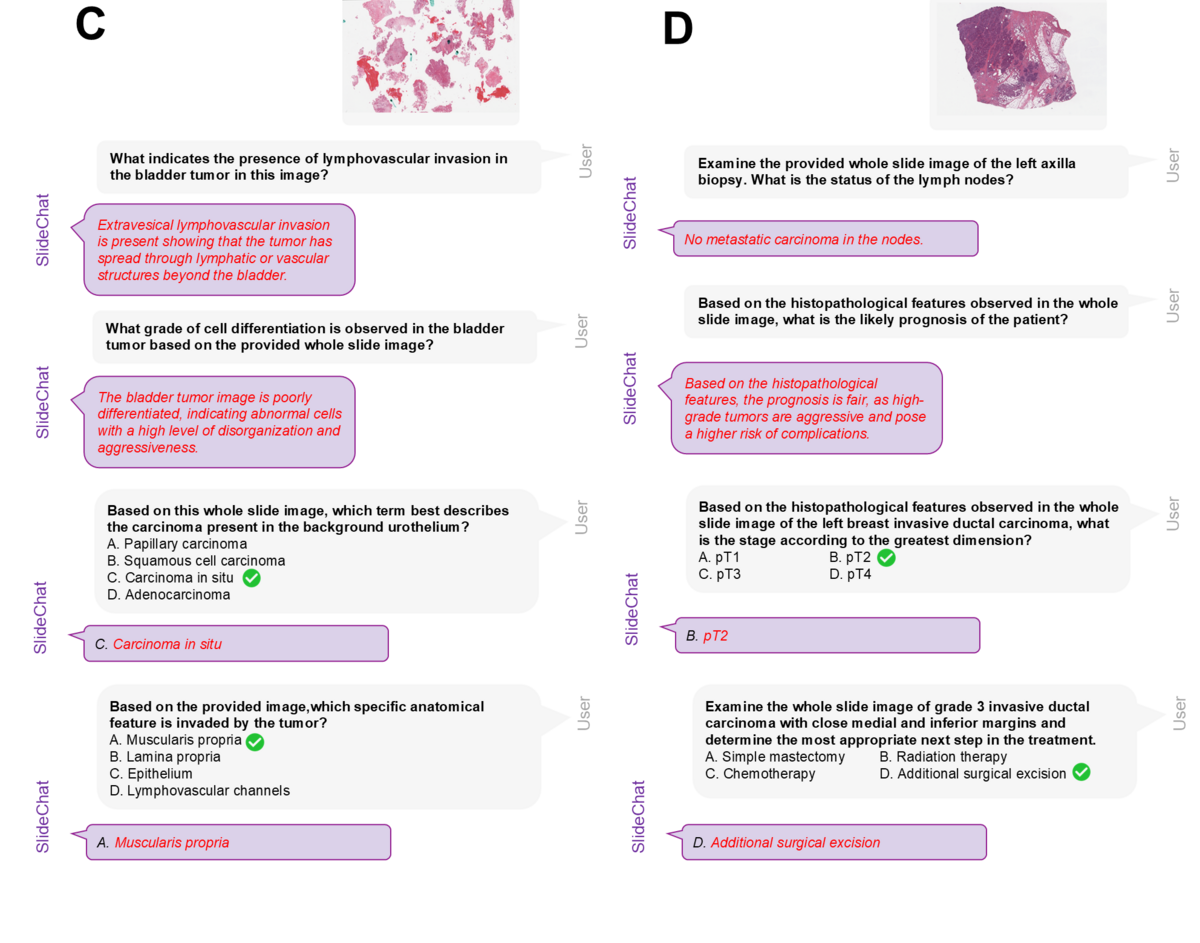
SlideChat supports multi-turn conversational reasoning over complete whole-slide images, enabling pathologists to ask follow-up questions about specific findings. Clinical examples include bladder tumor analysis (identifying lymphovascular invasion, cell differentiation grade, tumor type, anatomical invasion) and breast carcinoma assessment (lymph node metastasis status, prognosis prediction, staging by greatest dimension, treatment recommendation). The model provides clinically grounded, contextually accurate answers drawing on both local cellular detail and global tissue architecture — capabilities fundamentally unavailable to patch-only systems.
SlideChat represents a landmark advance in computational pathology — the first system to achieve genuine gigapixel whole-slide image understanding through conversational AI. By combining patch-level cellular precision with slide-level contextual reasoning via LongNet, and grounding training on the large-scale SlideInstruction dataset, SlideChat bridges the critical gap between patch-based analysis and clinically meaningful whole-slide interpretation. Its CVPR 2025 acceptance and SOTA results on 18/22 benchmarks, together with the full open-source release of model, dataset, and benchmark, establish a new foundation for AI-assisted pathology diagnosis, research, and education.
Key Contributions
- SlideInstruction: Largest WSI instruction-following dataset — 4.2K WSI-caption pairs and 176K VQA pairs across 13 clinical categories, curated from 4,915 TCGA pathology reports via GPT-4.
- SlideChat: First vision-language assistant for gigapixel WSI understanding — CONCH patch encoder + LongNet slide encoder + LLM, achieving SOTA on 18/22 tasks (CVPR 2025).
- SlideBench: Comprehensive WSI multi-modal benchmark spanning captioning and VQA across TCGA (1,494 → 3,176 samples, 10 → 31 cancer types), BCNB, CPTAC, and HISTAI subsets, all expert-validated.
- Open-source release: Full release of SlideChat model weights, SlideInstruction dataset, and SlideBench evaluation framework to accelerate computational pathology research.
Authors
Ying Chen*, Guoan Wang*, Yuanfeng Ji*†, Yanjun Li, Jin Ye, Tianbin Li, Ming Hu, Rongshan Yu, Yu Qiao, Junjun He†
* Equal contribution · † Corresponding authors
Shanghai AI Laboratory · Xiamen University · East China Normal University · Stanford University · Monash University