STU-Net: Scalable and Transferable Medical Image Segmentation Models
A family of scalable U-Net models ranging from 14M to 1.4B parameters, pre-trained on TotalSegmentator for universal medical image segmentation
🏆 MICCAI 2023 SPPIN Challenge — Champion
🥈 MICCAI 2023 AutoPET II Challenge — Runner-up (Highest DSC)
🥈 MICCAI 2023 BraTS2023 — Runner-up (+ two 3rd-place finishes)
🥉 FLARE 2023 — 3rd Place
Large-scale pre-trained models have transformed natural language processing and computer vision — yet medical image segmentation has remained dominated by small-scale models with only tens of millions of parameters. Scaling these models to higher orders of magnitude, and establishing whether larger models actually transfer better across clinical tasks, was an open question before STU-Net.
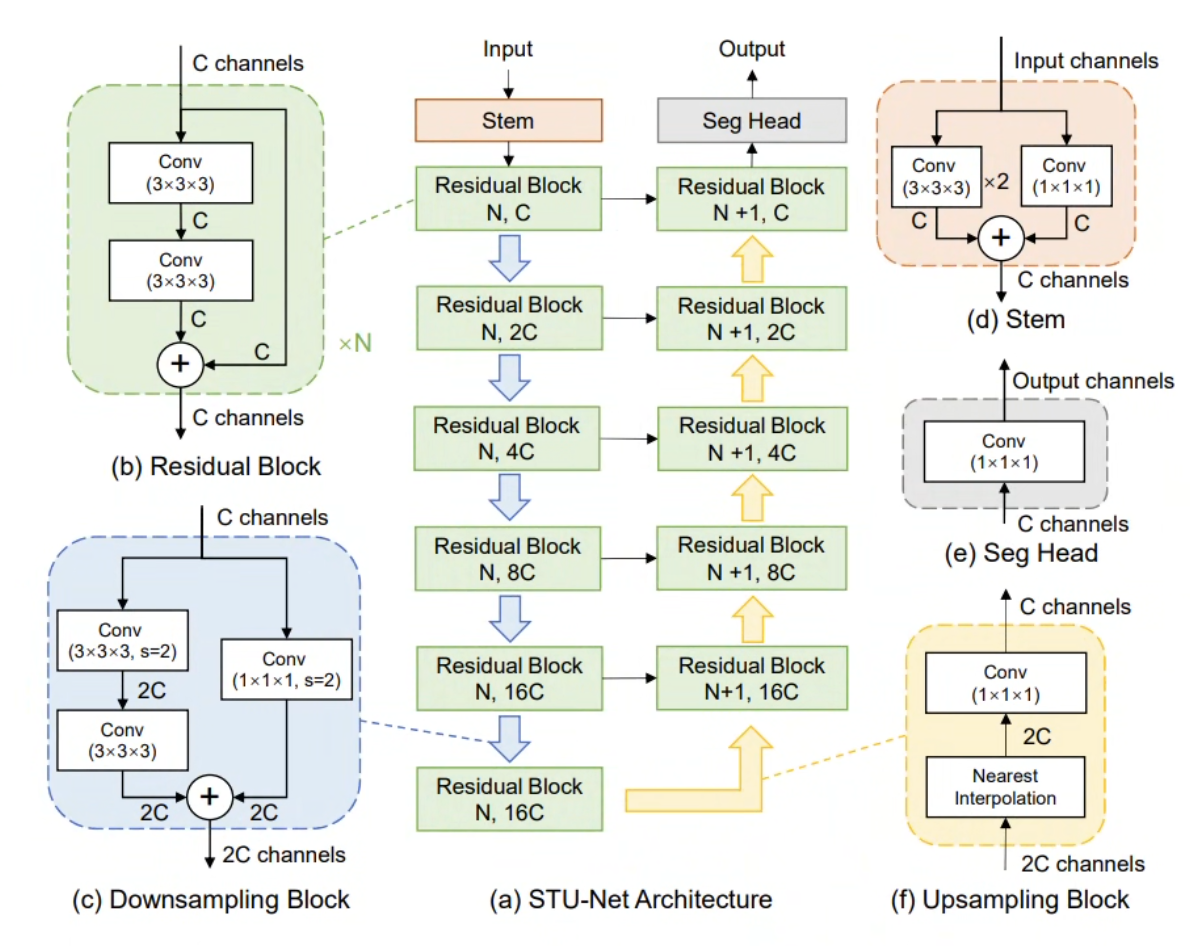
We designed a series of Scalable and Transferable U-Net (STU-Net) models with parameter counts ranging from 14M (STU-Net-S) to 1.4B (STU-Net-H). STU-Net-H is the largest medical image segmentation model to date. All variants are built on the nnU-Net framework with key architectural refinements: residual connections for deep scalability, and weight-free interpolation-based upsampling to eliminate the weight-mismatch problem during transfer learning.
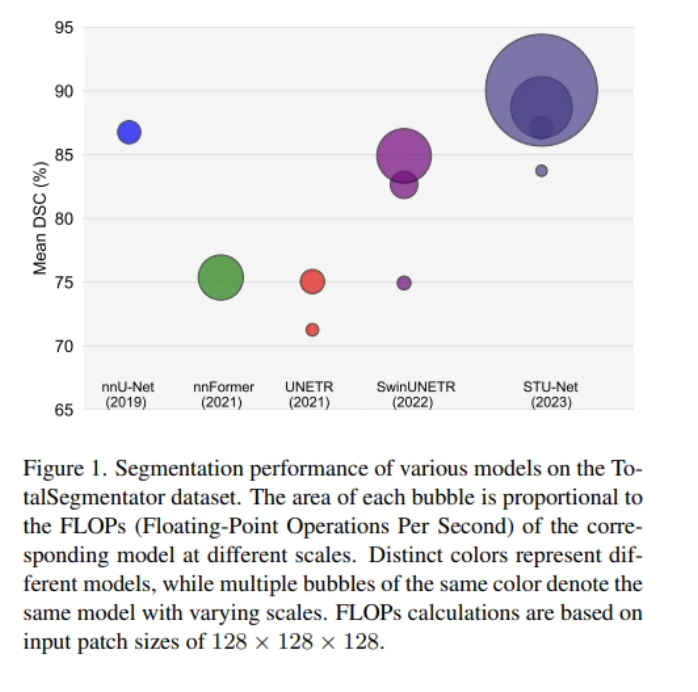
Pre-trained on TotalSegmentator — 1,204 CT volumes covering 104 anatomical structures — STU-Net demonstrates that scaling consistently improves segmentation accuracy. On the TotalSegmentator benchmark, STU-Net-H achieves 90.06% mean DSC, outperforming all CNN and Transformer competitors. Its transferability extends to 14 downstream datasets for direct inference and 3 datasets for fine-tuning, covering diverse modalities (CT, MRI, PET) and segmentation targets.
🌟 Core Highlights
01 — Scalability: Four Model Sizes from 14M to 1.4B Parameters
STU-Net comes in four sizes — S (14.6M), B (58.3M), L (440M), and H (1.46B parameters). The scaling strategy jointly increases network depth and width, which outperforms scaling either dimension alone. STU-Net-B already surpasses nnU-Net by 0.36% and SwinUNETR-B by 4.48% in mean DSC on TotalSegmentator. STU-Net-H achieves 90.06% mean DSC — the highest ever reported on this benchmark.
The architectural refinements make scaling possible: residual connections in each block prevent gradient diffusion in very deep networks, while the fixed 6-stage, isotropic-kernel configuration ensures that pre-trained weights are reusable across tasks without shape mismatch.
02 — Transferability: Strong Zero-Shot and Fine-tuned Performance Across 17 Datasets
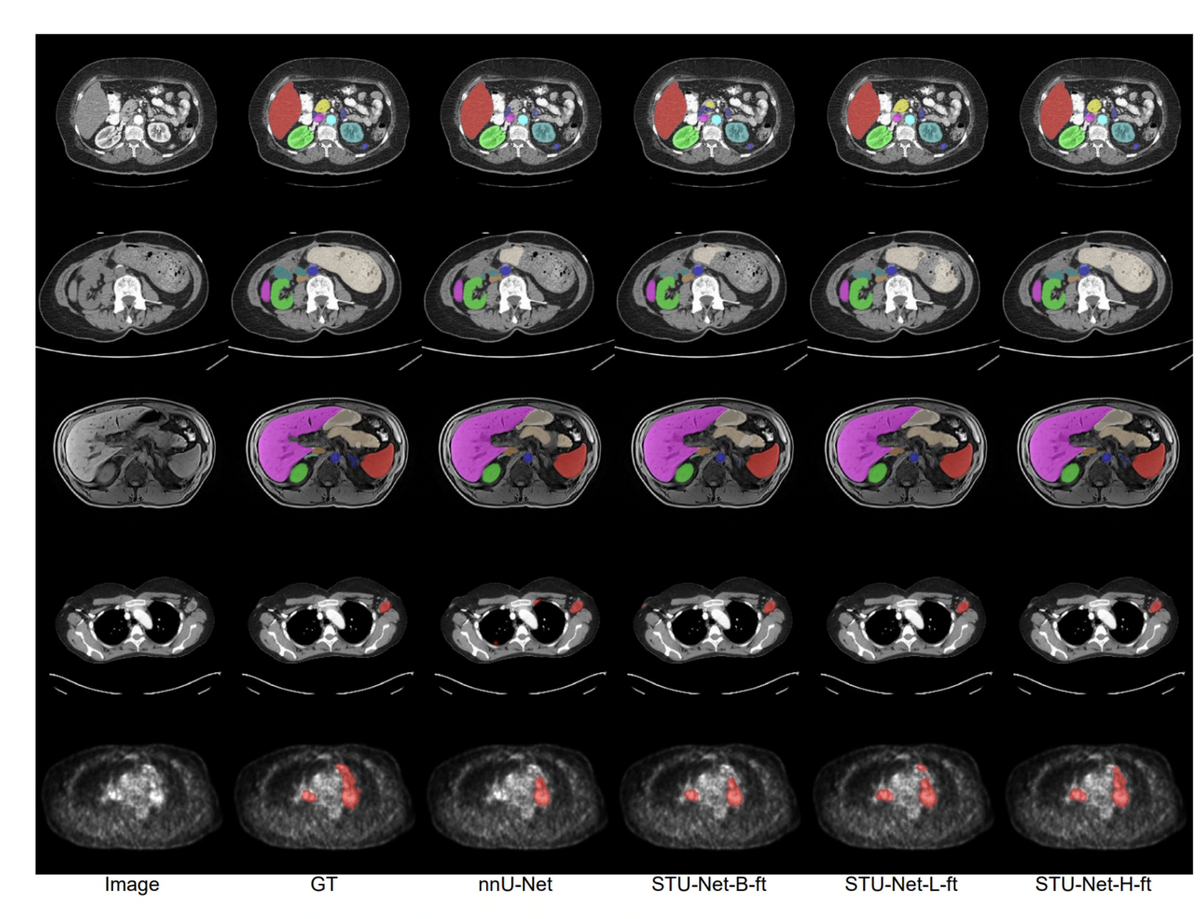
Pre-trained on TotalSegmentator, STU-Net can directly infer on 14 downstream CT datasets containing a subset of the 104 pre-training classes — no additional training required. Across these 14 datasets (2,494 cases total), STU-Net-H achieves 84.02% mean DSC vs. nnU-Net's 76.37%, a gain of 7.65%.
For fine-tuning on three challenging downstream datasets — FLARE22, AMOS22 (CT + MR), and AutoPET22 (CT + PET) — STU-Net-H-ft reaches 80.69% mean DSC vs. nnU-Net's 77.06%. Remarkably, fine-tuning on non-CT modalities (MRI, PET) also benefits from CT pre-training, suggesting the model captures fundamental anatomical structures that generalise beyond modality-specific features.
03 — State-of-the-Art Performance on TotalSegmentator
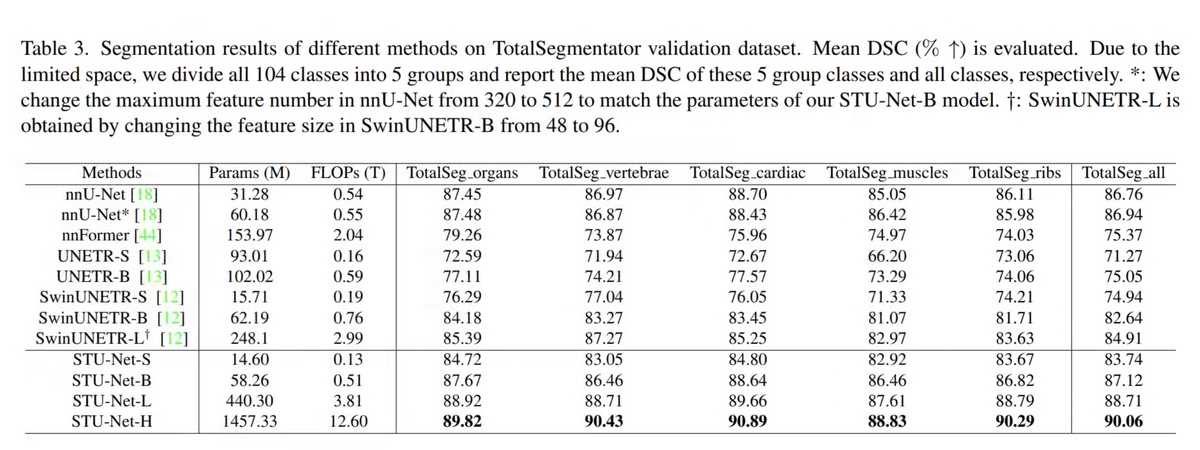
On the TotalSegmentator validation set — the largest publicly available CT segmentation benchmark with 104 structure annotations across organs, vertebrae, cardiac structures, muscles, and ribs — STU-Net-H achieves 90.06% mean DSC. This surpasses the previous best CNN model (nnU-Net: 86.76%) by +3.3% and the best Transformer model (SwinUNETR-B: 82.64%) by +7.4%.
The improvement is consistent across all five anatomical sub-groups, with the most notable gains in vertebrae (nnU-Net: 86.97% → STU-Net-H: 90.43%) and ribs (nnU-Net: 86.11% → STU-Net-H: 90.29%). This demonstrates that scaling genuinely improves comprehensiveness, not just overall average performance.
04 — Universal Models Surpass Specialist Models at Scale
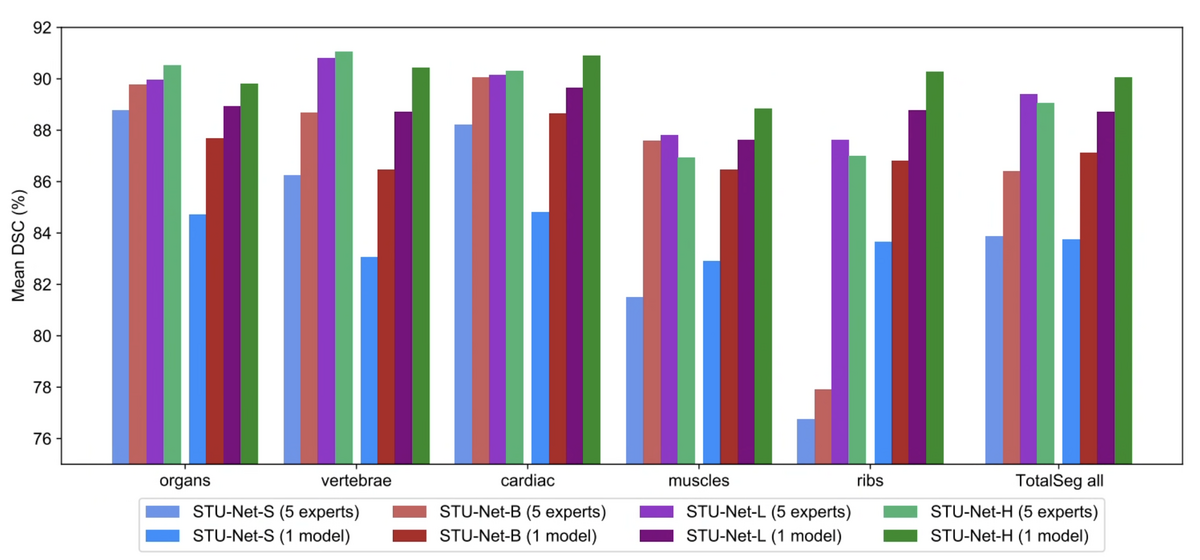
A long-standing assumption in medical image segmentation is that specialist models — trained on a single category group — outperform universal models handling all classes simultaneously. STU-Net challenges this assumption.
We trained five specialist models (organs, vertebrae, cardiac, muscles, ribs) and compared them against a single universal STU-Net trained on all 104 classes. At the STU-Net-H scale (1.4B parameters), the universal model achieves 90.06% overall mean DSC, surpassing the best specialist ensemble (89.07%). This suggests that at sufficient scale, a single unified model can simultaneously master all segmentation targets — a key step toward a true medical segmentation foundation model.
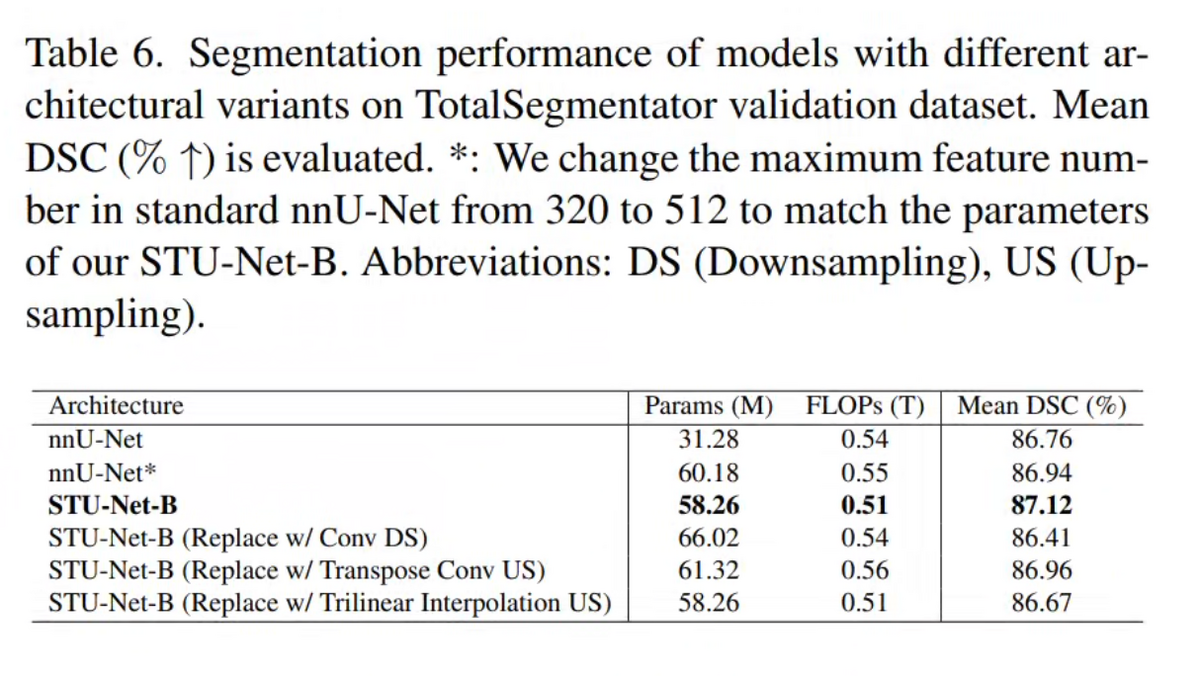
05 — Model Variants: Jointly Scaling Depth and Width
The four STU-Net variants are defined by systematic joint scaling of encoder depth and channel width: S (14.6M params, 12.8B FLOPs), B (58.3M, 60.9B), L (440M, 416B), and H (1.46B, 1,623B). Empirical ablations show that depth-only or width-only scaling yields diminishing returns compared to balanced joint scaling. Despite the 100× parameter gap between S and H, all variants share an identical 6-stage encoder-decoder topology and isotropic kernel configuration — this design constraint enables true weight transferability without shape-mismatch adapters.
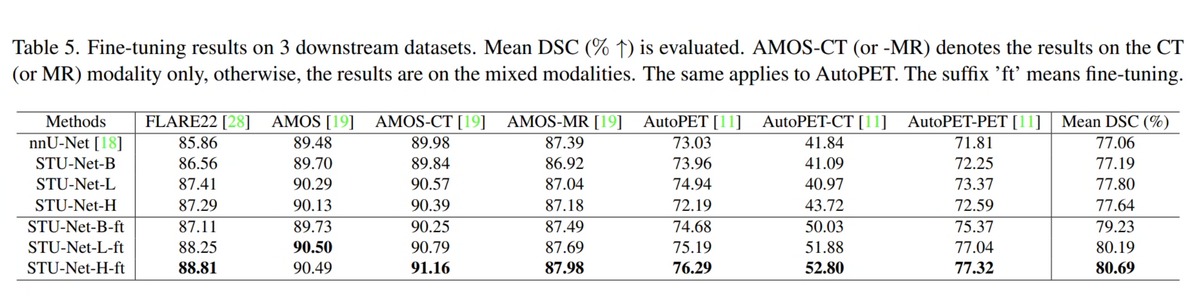
06 — Cross-Modality Transfer: Fine-Tuning on Downstream Datasets
On three challenging fine-tuning benchmarks — FLARE22 (13 abdominal organs), AMOS22 (CT + MRI, 15 organs), and AutoPET22 (CT + PET lesion segmentation) — STU-Net-H fine-tuned from pre-trained weights consistently outperforms nnU-Net fine-tuned from random initialization. The cross-modality transfer result is particularly notable: STU-Net-H pre-trained on CT only, when fine-tuned on AMOS-MRI and AutoPET-PET, achieves higher DSC than nnU-Net trained from scratch on those modalities — suggesting the pre-trained weights encode modality-agnostic anatomical priors.
STU-Net establishes that the scaling laws observed in natural language and computer vision do apply to 3D medical image segmentation. With 1.4B parameters and strong transferability across 17 datasets spanning CT, MRI, and PET modalities, STU-Net-H represents the current frontier of universal medical segmentation. It is a foundation model building block for Medical Artificial General Intelligence (MedAGI).
Key Contributions
- Designed STU-Net-S/B/L/H — scaling from 14M to 1.4B parameters; STU-Net-H is the largest medical image segmentation model to date.
- Demonstrated clear scaling laws: larger models trained on TotalSegmentator consistently achieve higher DSC on both the pre-training benchmark and 14 downstream transfer datasets.
- Refined nnU-Net architecture with residual blocks and weight-free interpolation upsampling for true cross-task weight transferability.
- Won championship at MICCAI 2023 ATLAS and SPPIN challenges; runner-up at AutoPET II; multiple top-3 finishes at BraTS2023 and FLARE2023.
Authors
Ziyan Huang, Haoyu Wang, Zhongying Deng, Jin Ye, Yanzhou Su, Hui Sun, Junjun He, Yun Gu, Lixu Gu, Shaoting Zhang, Yu Qiao
arXiv 2023